R Tutorial¶
The purpose of this tutorial is to walk the new user through examples demonstrating the use of H2O through R. The objective is to learn the basic syntax of H2O, including importing and parsing files, specifying a model, and obtaining model output.
Those who have never used H2O before should see the quick start guide for additional instructions on how to run H2O. Additionally, users who are using H2O through R for the first time will need to install the R package, available in our download package at: http://0xdata.com/downloadtable/.
It is highly recommended that users review the getting started section before proceeding to other sections for examples. At a minimum users should run the following commands before running any other examples, as the H2O library and H2O object (named “localH2O” in examples is required in R for most examples to work.
library(h2o)
localH2O = h2o.init(ip = "localhost", port = 54321, startH2O = TRUE)
In this tutorial you can find information on: Getting Started (establishing a connection between H2O and R), Importing Data, Data Manipulation, Running Models, Obtaining Predictions, Other Useful Functions.
Getting Started¶
Installing and Starting H2O
Before beginning, be sure to have an instance of H2O running. Additionally, users who are using H2O for the first time can find help for installing the R package at http://docs.0xdata.com/Ruser/Rh2opackage.html.
Step 1
Call the H2O package, and initialize H2O in R. Note that an object “localh2o” is created. Assigning the H2O initialization to an object is important, because the connection will be used later to tell R where to send data sets, model specification, and where to find results.
library(h2o)
localH2O <- h2o.init(ip = "localhost", port = 54321)
Users may see a response message in R indicating that the instance of H2O running is a different version than that of the corresponding H2O R package. This error will look similar to the picture below. If you get this error, it can be resolved by downloading the correct version of H2O from http://0xdata.com/downloadtable/. Users should follow the installation instructions on the download page.
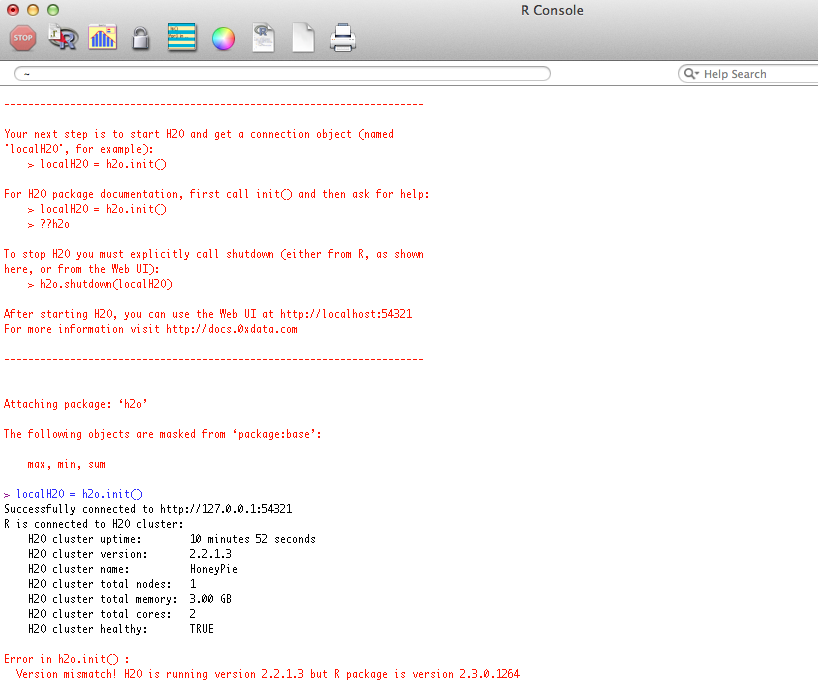
Cluster Info
Used to check that the H2O instance is running and healthy.
library(h2o)
localH2O = h2o.init(ip = "localhost", port = 54321, startH2O = TRUE)
h2o.checkClient(localH2O)
Importing Data¶
Import File
Use this call when importing a data set that exists in a single file.
irisPath = system.file("extdata", "iris.csv", package="h2o")
iris.hex = h2o.importFile(localH2O, path = irisPath, key = "iris.hex")
summary(iris.hex)
Import Folder
Use this call when importing a data set that exists in multiple files.
myPath = system.file("extdata", "prostate_folder", package = "h2o")
prostate_all.hex = h2o.importFolder(localH2O, path = myPath)
class(prostate_all.hex)
summary(prostate_all.hex)
prostate_all.fv = h2o.importFolder(localH2O, path = myPath, version = 2)
Import URL
Use this call when data are stored at a website accessible from the machine on which the instance of H2O is running.
prostate.hex = h2o.importURL(localH2O, path = paste("https://raw.github.com",
"0xdata/h2o/master/smalldata/logreg/prostate.csv", sep = "/"), key = "prostate.hex")
class(prostate.hex)
summary(prostate.hex)
Data Manipulation and Description¶
Any Factor
Used to determine if any column in a data set is a factor.
irisPath = system.file("extdata", "iris_wheader.csv", package="h2o")
iris.hex = h2o.importFile(localH2O, path = irisPath)
h2o.anyFactor(iris.hex)
As Data Frame
Used to convert an H2O parsed data object into an R data frame (which can subsequently be manipulated using R calls). While this is frequently useful, as.data.frame should be used with care when converting H2O Parsed Data objects. Data sets that are easily and quickly handled by H2O are often too large to be treated equivalently well in R.
prosPath <- system.file("extdata", "prostate.csv", package="h2o")
prostate.hex = h2o.importFile(localH2O, path = prosPath)
prostate.data.frame<- as.data.frame(prostate.hex)
summary(prostate.data.frame)
head(prostate.data.frame)
As Factor
Used to convert an integer into a non-ordered factor (alternatively called an enum or categorical).
prosPath = system.file("extdata", "prostate.csv", package="h2o")
prostate.hex = h2o.importFile(localH2O, path = prosPath)
prostate.hex[,4] = as.factor(prostate.hex[,4])
summary(prostate.hex)
As H2O
Used to pass a data frame from inside of the R environment to the H2O instance.
data(iris)
summary(iris)
iris.r <- iris
iris.h2o <- as.h2o(localH2O, iris.r, key="iris.h2o")
class(iris.h2o)
Assign H2O
Used to create an hex key on the server where H2O is running for data sets manipulated in R. For instance, in the example below, the prostate data set was uploaded to the H2O instance, and was manipulated to remove outliers. Saving the new data set on the H2O server so that it can be subsequently be analyzed with H2O without overwriting the original data set relies on h2o.assign.
prosPath = system.file("extdata", "prostate.csv", package="h2o")
prostate.hex = h2o.importFile(localH2O, path = prosPath)
prostate.qs = quantile(prostate.hex$PSA)
PSA.outliers = prostate.hex[prostate.hex$PSA <= prostate.qs[2] | prostate.hex$PSA >= prostate.qs[10],]
PSA.outliers = h2o.assign(PSA.outliers, "PSA.outliers")
nrow(prostate.hex)
nrow(PSA.outliers)
Colnames
Used to obtain a list of the column names in a data set.
irisPath = system.file("extdata", "iris.csv", package="h2o")
iris.hex = h2o.importFile(localH2O, path = irisPath, key = "iris.hex")
summary(iris.hex)
colnames(iris.hex)
Extremes
Used to obtain the maximum and minimum values in real valued columns.
ausPath = system.file("extdata", "australia.csv", package="h2o")
australia.hex = h2o.importFile(localH2O, path = ausPath, key = "australia.hex")
min(australia.hex)
min(c(-1, 0.5, 0.2), FALSE, australia.hex[,1:4])
Quantiles
Used to request quantiles for an H2O parsed data set. When requested for a full parsed data set quantiles() returns a matrix displaying quantile information for all numeric columns in the data set.
prosPath = system.file("extdata", "prostate.csv", package="h2o")
prostate.hex = h2o.importFile(localH2O, path = prosPath)
quantile(prostate.hex)
Summary
Used to generate an R like summary for each of the columns of a data set. For continuous reals this produces a summary that includes information on quartiles, min, max and mean. For factors this produces information on counts of elements within each factor level. For information on the Summary algorithm see Summary
prosPath = system.file("extdata", "prostate.csv", package="h2o")
prostate.hex = h2o.importFile(localH2O, path = prosPath)
summary(prostate.hex)
summary(prostate.hex$GLEASON)
summary(prostate.hex[,4:6])
H2O Table
Used to summarize information in data. Note that because H2O handles such large data sets,
it is possible for users to generate tables that are larger that R’s capacity. To minimize this risk and allow users to work uninterrruped, h2o.table is called inside of a call for head() or tail(). Within head() and tail() users can explicity specify the number of rows in the table to return.
head(h2o.table(prostate.hex[,3]))
head(h2o.table(prostate.hex[,c(3,4)]))
Test Train Split and generating Random Numbers
Runif is used to append a column of random numbers to an H2O data frame and facilitate creating test/ train splits of data for analysis and validation in H2O.
prosPath = system.file("extdata", "prostate.csv", package="h2o")
prostate.hex = h2o.importFile(localH2O, path = prosPath, key = "prostate.hex")
s = h2o.runif(prostate.hex)
summary(s)
prostate.train = prostate.hex[s <= 0.8,]
prostate.train = h2o.assign(prostate.train, "prostate.train")
prostate.test = prostate.hex[s > 0.8,]
prostate.test = h2o.assign(prostate.test, "prostate.test")
nrow(prostate.train) + nrow(prostate.test)
Running Models¶
GBM
Gradient Boosted Models. For information on the GBM algorithm see Gradient Boosted Regression and Classification
ausPath = system.file("extdata", "australia.csv", package="h2o")
australia.hex = h2o.importFile(localH2O, path = ausPath)
independent <- c("premax", "salmax","minairtemp", "maxairtemp",
"maxsst", "maxsoilmoist", "Max_czcs")
dependent <- "runoffnew"
h2o.gbm(y = dependent, x = independent, data = australia.hex,
n.trees = 10, interaction.depth = 3,
n.minobsinnode = 2, shrinkage = 0.2, distribution= "gaussian")
Run multinomial classification GBM on abalone data
h2o.gbm(y = dependent, x = independent, data = australia.hex, n.trees
= 15, interaction.depth = 5,
n.minobsinnode = 2, shrinkage = 0.01, distribution= "multinomial")
Generalized Linear Models
Generalized linear models, which are used to develop linear models for exponential distributions. Regularization can be applied. For information on the GBM algorithm see Generalized Linear Model (GLM)
prostate.hex = h2o.importURL.VA(localH2O, path =
"https://raw.github.com/0xdata/h2o/master/smalldata/logreg/prostate.csv",
key = "prostate.hex")
h2o.glm(y = "CAPSULE", x = c("AGE","RACE","PSA","DCAPS"), data =
prostate.hex, family = "binomial", nfolds = 10, alpha = 0.5)
myX = setdiff(colnames(prostate.hex), c("ID", "DPROS", "DCAPS", "VOL"))
h2o.glm(y = "VOL", x = myX, data = prostate.hex, family = "gaussian", nfolds = 5, alpha = 0.1)
K-Means
K means is a clustering algorithm that allows users to characterize data. This algorithm does not rely on a dependent variable. For information on the K-Means algorithm see K-Means
prosPath = system.file("extdata", "prostate.csv", package="h2o")
prostate.hex = h2o.importFile(localH2O, path = prosPath)
h2o.kmeans(data = prostate.hex, centers = 10, cols = c("AGE", "RACE", "VOL", "GLEASON"))
covPath = system.file("extdata", "covtype.csv", package="h2o")
covtype.hex = h2o.importFile(localH2O, path = covPath)
covtype.km = h2o.kmeans(data = covtype.hex, centers = 5, cols = c(1, 2, 3))
print(covtype.km)
Principal Components Analysis
Principal Components Analysis maps a set of variables onto a subspace via linear transformations. PCA is the first step in Principal Components Regression. For more information on PCA see Principal Components Analysis.
ausPath = system.file("extdata", "australia.csv", package="h2o")
australia.hex = h2o.importFile(localH2O, path = ausPath)
australia.pca = h2o.prcomp(data = australia.hex, standardize = TRUE)
print(australia.pca)
summary(australia.pca)
Principal Components Regression
PCR is an algorithm that allows users to map a set of variables to a new set of linearly independent variables. The new set of variables are linearly independent linear combinations of the original variables and exist in a subspace of lower dimension. This transformation is then prepended to a regression model, often improving results. For more information on PCA see Principal Components Analysis.
prostate.hex = h2o.importFile(localH2O,
path =
"https://raw.github.com/0xdata/h2o/master/smalldata/logreg/prostate.csv",
key = "prostate.hex")
h2o.pcr(x = c("AGE","RACE","PSA","DCAPS"), y = "CAPSULE", data =
prostate.hex, family = "binomial",
nfolds = 10, alpha = 0.5, ncomp = 3)
Obtaining Predictions¶
Predict
Used to apply an H2O model to a holdout set to obtain predictions based on model results. In the examples below models are first generated, and then the predictions for that model are obtained.
prostate.hex = h2o.importURL.VA(localH2O,
path =
"https://raw.github.com/0xdata/h2o/master/smalldata/logreg/prostate.csv",
key = "prostate.hex")
prostate.glm = h2o.glm(y = "CAPSULE", x =
c("AGE","RACE","PSA","DCAPS"), data = prostate.hex,
family = "binomial", nfolds = 10, alpha = 0.5)
prostate.fit = h2o.predict(object = prostate.glm, newdata = prostate.hex)
summary(prostate.fit)
covPath = system.file("extdata", "covtype.csv", package="h2o")
covtype.hex = h2o.importFile(localH2O, path = covPath)
covtype.km = h2o.kmeans(data = covtype.hex, centers = 5, cols = c(1, 2, 3))
covtype.clusters = h2o.predict(object = covtype.km, newdata = covtype.hex)
Other Useful Functions¶
List all H2O Objects
Used to generate a list of all H2O objects that have been generated during a work session, along with each objects byte size.
prostate.hex = h2o.importFile(localH2O, path = prosPath, key = "prostate.hex")
s = runif(nrow(prostate.hex))
prostate.train = prostate.hex[s <= 0.8,]
prostate.train = h2o.assign(prostate.train, "prostate.train")
h2o.ls(localH2O)
Remove an H2O object from the server where H2O is running
Users may wish to remove an H2O object on the server that is associated with an object in the R environment. Recommended behavior is to also remove the object in the R environment.
localH2O = h2o.init()
prosPath = system.file("extdata", "prostate.csv", package="h2o")
prostate.hex = h2o.importFile(localH2O, path = prosPath, key = "prostate.hex")
s = runif(nrow(prostate.hex))
prostate.train = prostate.hex[s <= 0.8,]
prostate.train = h2o.assign(prostate.train, "prostate.train")
h2o.ls(localH2O)
h2o.rm(object= localH2O, keys= "Last.value.0")
h2o.ls(localH2O)