Welcome to H2O 3.0
- New Users
- Experienced Users
- Enterprise Users
- Sparkling Water Users
- Python Users
- R Users
- API Users
- Java Users
- Developers
Welcome to the H2O documentation site! Depending on your area of interest, select a learning path from the links above.
We’re glad you’re interested in learning more about H2O - if you have any questions, please email them to support@h2o.ai or post them on our Google groups website, h2ostream.
Note: To join our Google group on h2ostream, you need a Google account (such as Gmail or Google+). On the h2ostream page, click the Join group button, then click the New Topic button to post a new message. You don’t need to request or leave a message to join - you should be added to the group automatically.
We welcome your feedback! Please let us know if you have any questions or comments about this site by emailing us at support@h2o.ai.
New Users
If you’re just getting started with H2O, here are some links to help you learn more:
Downloads page: First things first - download a copy of H2O here by selecting a build under “Download H2O” (the “Bleeding Edge” build contains the latest changes, while the latest alpha release represents a more stable build), then use the installation instruction tabs to install H2O on your client of choice (standalone, R, Python, Hadoop, or Maven) .
For first-time users, we recommend downloading the latest alpha release and the default standalone option (the first tab) as the installation method. Make sure to install Java if it is not already installed.
The following video provides step-by-step instructions on how to install and run H2O:
Tutorials: To see a step-by-step example of our algorithms in action, select a model type from the following list:
Getting Started with Flow: This document describes our new intuitive web interface, Flow. This interface is similar to IPython notebooks, and allows you to create a visual workflow to share with others.
Launch from the command line: This document describes some of the additional options that you can configure when launching H2O (for example, to specify a different directory for saved Flow data, allocate more memory, or use a flatfile for quick configuration of a cluster).
Algorithms: This document describes the science behind our algorithms and provides a detailed, per-algo view of each model type.
Experienced Users
If you’ve used previous versions of H2O, the following links will help guide you through the process of upgrading to H2O 3.0.
Migration Guide: This document provides a comprehensive guide to assist users in upgrading to H2O 3.0. It gives an overview of the changes to the algorithms and the web UI introduced in this version and describes the benefits of upgrading for users of R, APIs, and Java.
Porting R Scripts: This document is designed to assist users who have created R scripts using previous versions of H2O. Due to the many improvements in R, scripts created using previous versions of H2O need some revision to work with H2O 3.0. This document provides a side-by-side comparison of the changes in R for each algorithm, as well as overall structural enhancements R users should be aware of, and provides a link to a tool that assists users in upgrading their scripts.
Recent Changes: This document describes the most recent changes in the latest build of H2O. It lists new features, enhancements (including changed parameter default values), and bug fixes for each release, organized by sub-categories such as Python, R, and Web UI.
H2O Classic vs H2O 3.0: This document presents a side-by-side comparison of H2O 3.0 and the previous version of H2O. It compares and contrasts the features, capabilities, and supported algorithms between the versions. If you’d like to learn more about the benefits of upgrading, this is a great source of information.
Algorithms Roadmap: This document outlines our currently implemented features and describes which features are planned for future software versions. If you’d like to know what’s up next for H2O, this is the place to go.
Contributing code: If you’re interested in contributing code to H2O, we appreciate your assistance! This document describes how to access our list of Jiras that are suggested tasks for contributors and how to contact us.
Enterprise Users
If you’re considering using H2O in an enterprise environment, you’ll be happy to know that H2O supports many popular scalable computing solutions, such as Hadoop and EC2 (AWS). For more information, refer to the following links.
The following video provides step-by-step instructions on how to start H2O on Hadoop:
How to Pass S3 Credentials to H2O: This document describes the necessary step of passing your S3 credentials to H2O so that H2O can be used with AWS, as well as how to run H2O on an EC2 cluster.
Running H2O on Hadoop: This document describes how to run H2O on Hadoop.
Sparkling Water Users
Sparkling Water is a gradle project with the following submodules:
- Core: Implementation of H2OContext, H2ORDD, and all technical integration code
- Examples: Application, demos, examples
- ML: Implementation of MLLib pipelines for H2O algorithms
- Assembly: Creates “fatJar” composed of all other modules
The best way to get started is to modify the core module or create a new module, which extends a project.
Users of our Spark-compatible solution, Sparkling Water, should be aware that Sparkling Water is only supported with the latest version of H2O. For more information about Sparkling Water, refer to the following links.
Getting Started with Sparkling Water
The following video provides step-by-step instructions on how to start H2O using Sparkling Water:
Download Sparkling Water: Go here to download Sparkling Water.
Sparkling Water Development Documentation: Read this document first to get started with Sparkling Water.
Launch on Hadoop and Import from HDFS: Go here to learn how to start Sparkling Water on Hadoop.
Sparkling Water Tutorials: Go here for demos and examples.
Sparkling Water K-means Tutorial: Go here to view a demo that uses Scala to create a K-means model.
Sparkling Water GBM Tutorial: Go here to view a demo that uses Scala to create a GBM model.
Sparkling Water on YARN: Follow these instructions to run Sparkling Water on a YARN cluster.
Building Applications on top of H2O: This short tutorial describes project building and demonstrates the capabilities of Sparkling Water using Spark Shell to build a Deep Learning model.
Sparkling Water FAQ: This FAQ provides answers to many common questions about Sparkling Water.
Sparkling Water Blog Posts
Sparkling Water Meetup Slide Decks
Python Users
Pythonistas will be glad to know that H2O now provides support for this popular programming language. Python users can also use H2O with IPython notebooks. For more information, refer to the following links.
The following video provides step-by-step instructions on how to start H2O using Python:
Python readme: This document describes how to setup and install the prerequisites for using Python with H2O.
Python docs: This document represents the definitive guide to using Python with H2O.
R Users
Don’t worry, R users - we still provide R support in the latest version of H2O, just as before. The R components of H2O have been cleaned up, simplified, and standardized, so the command format is easier and more intuitive. Due to these improvements, be aware that any scripts created with previous versions of H2O will need some revision to be compatible with the latest version.
We have provided the following helpful resources to assist R users in upgrading to the latest version, including a document that outlines the differences between versions and a tool that reviews scripts for deprecated or renamed parameters.
The following video provides step-by-step instructions on how to start H2O in R:
R User Documentation: This document contains all commands in the H2O package for R, including examples and arguments. It represents the definitive guide to using H2O in R.
Porting R Scripts: This document is designed to assist users who have created R scripts using previous versions of H2O. Due to the many improvements in R, scripts created using previous versions of H2O will not work. This document provides a side-by-side comparison of the changes in R for each algorithm, as well as overall structural enhancements R users should be aware of, and provides a link to a tool that assists users in upgrading their scripts.
API Users
API users will be happy to know that the APIs have been more thoroughly documented in the latest release of H2O and additional capabilities (such as exporting weights and biases for Deep Learning models) have been added.
REST APIs are generated immediately out of the code, allowing users to implement machine learning in many ways. For example, REST APIs could be used to call a model created by sensor data and to set up auto-alerts if the sensor data falls below a specified threshold.
REST API Reference: This document represents the definitive guide to the H2O REST API.
REST API Schema Reference: This document represents the definitive guide to the H2O REST API schemas.
Java Users
For Java developers, the following resources will help you create your own custom app that uses H2O.
H2O Core Java Developer Documentation: The definitive Java API guide for the core components of H2O.
H2O Algos Java Developer Documentation: The definitive Java API guide for the algorithms used by H2O.
Developers
If you’re looking to use H2O to help you develop your own apps, the following links will provide helpful references.
For IDEA IntelliJ support, run gradle idea, then Import Project within IDEA and point it to the h2o-3 directory.
For JUnit tests to pass, you may need multiple H2O nodes. Create a “Run/Debug” configuration with the following parameters:
Type: Application
Main class: H2OApp
Use class path of module: h2o-app
After starting multiple “worker” node processes in addition to the JUnit test process, they will cloud up and run the multi-node JUnit tests.
Maven install: This page provides information on how to build a version of H2O that generates the correct IDE files.
apps.h2o.ai: Apps.h2o.ai is designed to support application developers via events, networking opportunities, and a new, dedicated website comprising developer kits and technical specs, news, and product spotlights.
H2O Project Templates: This page provides template info for projects created in Java, Scala, or Sparkling Water.
H2O Scala API Developer Documentation: The definitive Scala API guide for H2O.
Contributing code: If you’re interested in contributing code to H2O, we appreciate your assistance! This document describes how to access our list of Jiras that contributors can work on and how to contact us.
Downloading H2O
To download H2O, go to our downloads page. Select a build type (bleeding edge or latest alpha), then select an installation method (standalone, R, Python, Hadoop, or Maven) by clicking the tabs at the top of the page. Follow the instructions in the tab to install H2O.
Starting H2O …
There are a variety of ways to start H2O, depending on which client you would like to use.
… From R
To use H2O in R, follow the instructions on the download page.
… From Python
To use H2O in Python, follow the instructions on the download page.
… On Spark
To use H2O on Spark, follow the instructions on the Sparkling Water download page.
… From the Cmd Line
You can use Terminal (OS X) or the Command Prompt (Windows) to launch H2O 3.0. When you launch from the command line, you can include additional instructions to H2O 3.0, such as how many nodes to launch, how much memory to allocate for each node, assign names to the nodes in the cloud, and more.
There are two different argument types:
- JVM arguments
- H2O arguments
The arguments use the following format: java <JVM Options> -jar h2o.jar <H2O Options>.
JVM Options
-version: Display Java version info.-Xmx<Heap Size>: To set the total heap size for an H2O node, configure the memory allocation option-Xmx. By default, this option is set to 1 Gb (-Xmx1g). When launching nodes, we recommend allocating a total of four times the memory of your data.
Note: Do not try to launch H2O with more memory than you have available.
H2O Options
hor-help: Display this information in the command line output.-name <H2OCloudName>: Assign a name to the H2O instance in the cloud (where<H2OCloudName>is the name of the cloud. Nodes with the same cloud name will form an H2O cloud (also known as an H2O cluster).-flatfile <FileName>: Specify a flatfile of IP address for faster cloud formation (where<FileName>is the name of the flatfile.-ip <IPnodeAddress>: Specify an IP address other than the defaultlocalhostfor the node to use (where<IPnodeAddress>is the IP address).-port <#>: Specify a port number other than the default54321for the node to use (where<#>is the port number).-network <IPv4NetworkSpecification1>[,<IPv4NetworkSpecification2> ...]: Specify a range (where applicable) of IP addresses (where<IPv4NetworkSpecification1>represents the first interface,<IPv4NetworkSpecification2>represents the second, and so on). The IP address discovery code binds to the first interface that matches one of the networks in the comma-separated list. For example,10.1.2.0/24supports 256 possibilities.-ice_root <fileSystemPath>: Specify a directory for H2O to spill temporary data to disk (where<fileSystemPath>is the file path).-flow_dir <server-side or HDFS directory>: Specify a directory for saved flows. The default is/Users/h2o-<H2OUserName>/h2oflows(where<H2OUserName>is your user name).nthreads <#ofThreads>: Specify the maximum number of threads in the low-priority batch work queue (where<#ofThreads>is the number of threads). The default is 99.-client: Launch H2O node in client mode. This is used mostly for running Sparkling Water.
Cloud Formation Behavior
New H2O nodes join to form a cloud during launch. After a job has started on the cloud, it prevents new members from joining.
To start an H2O node with 4GB of memory and a default cloud name:
java -Xmx4g -jar h2o.jarTo start an H2O node with 6GB of memory and a specific cloud name:
java -Xmx6g -jar h2o.jar -name MyCloudTo start an H2O cloud with three 2GB nodes using the default cloud names:
java -Xmx2g -jar h2o.jar &java -Xmx2g -jar h2o.jar &java -Xmx2g -jar h2o.jar &
Wait for the INFO: Registered: # schemas in: #mS output before entering the above command again to add another node (the number for # will vary).
Flatfile Configuration
If you are configuring many nodes, it is faster and easier to use the -flatfile option, rather than -ip and -port.
To configure H2O on a multi-node cluster:
- Locate a set of hosts.
- Download the appropriate version of H2O for your environment.
- Verify that the same h2o.jar file is available on all hosts.
Create a flatfile (a plain text file with the IP and port numbers of the hosts). Use one entry per line. For example:
192.168.1.163:54321 192.168.1.164:54321- Copy the flatfile.txt to each node in the cluster.
Use the
-Xmxoption to specify the amount of memory for each node. The cluster’s memory capacity is the sum of all H2O nodes in the cluster.For example, if you create a cluster with four 20g nodes (by specifying
-Xmx20gfour times), H2O will have a total of 80 gigs of memory available.For best performance, we recommend sizing your cluster to be about four times the size of your data. To avoid swapping, the
-Xmxallocation must not exceed the physical memory on any node. Allocating the same amount of memory for all nodes is strongly recommended, as H2O works best with symmetric nodes.Note the optional
-ipand-portoptions specify the IP address and ports to use. The-ipoption is especially helpful for hosts with multiple network interfaces.java -Xmx20g -jar h2o.jar -flatfile flatfile.txt -port 54321The output will resemble the following:
04-20 16:14:00.253 192.168.1.70:54321 2754 main INFO: 1. Open a terminal and run 'ssh -L 55555:localhost:54321 H2O-3User@###.###.#.##' 04-20 16:14:00.253 192.168.1.70:54321 2754 main INFO: 2. Point your browser to http://localhost:55555 04-20 16:14:00.437 192.168.1.70:54321 2754 main INFO: Log dir: '/tmp/h2o-H2O-3User/h2ologs' 04-20 16:14:00.437 192.168.1.70:54321 2754 main INFO: Cur dir: '/Users/H2O-3User/h2o-3' 04-20 16:14:00.459 192.168.1.70:54321 2754 main INFO: HDFS subsystem successfully initialized 04-20 16:14:00.460 192.168.1.70:54321 2754 main INFO: S3 subsystem successfully initialized 04-20 16:14:00.460 192.168.1.70:54321 2754 main INFO: Flow dir: '/Users/H2O-3User/h2oflows' 04-20 16:14:00.475 192.168.1.70:54321 2754 main INFO: Cloud of size 1 formed [/192.168.1.70:54321]As you add more nodes to your cluster, the output is updated:
INFO WATER: Cloud of size 2 formed [/...]...Access the H2O 3.0 web UI (Flow) with your browser. Point your browser to the HTTP address specified in the output
Listening for HTTP and REST traffic on ....To check if the cloud is available, point to the url
http://<ip>:<port>/Cloud.json(an example of the JSON response is provided below). Wait forcloud_sizeto be the expected value and theconsensusfield to be true:{ ... "cloud_size": 2, "consensus": true, ... }
Manual Multi-node
Running H2O on a multi-node cluster allows you to use more memory for large-scale tasks (for example, creating models from huge datasets) than would be possible on a single node.
Locate a set of hosts that will be used to create your cluster. A host can be a server, an EC2 instance, or your laptop.
Download H2O, including the .jar file, by going to the H2O downloads page and choosing the appropriate version for your environment.
Verify the same h2o.jar file is available on each host in the multi-node cluster.
Create a flatfile.txt that contains an IP address and port number for each H2O instance. Use one entry per line. For example:
192.168.1.163:54321 192.168.1.164:54321A flat file listing the nodes is the easiest way to get multiple H2O nodes to find each other and form a cluster. Note that the
-flatfileoption tells one H2O node where to find the others. It is not a substitute for the-ipand-portspecification.Copy the flatfile.txt to each node in your cluster.
Use the
-Xmxoption in the Java command line to specify the amount of memory allocated to each H2O node. The cluster’s memory capacity is the sum of the memory available across all H2O nodes in the cluster.For example, if you create a cluster with four 20g nodes (by specifying
-Xmx20g), H2O will have a total of 80 gigs of memory available.For best performance, we recommend creating a cluster about four times the size of your data. However, to avoid memory swapping, the Xmx value must not be larger than the physical memory on any given node. We strongly recommend allocating the same amount of memory for all nodes, since H2O works best with symmetric nodes.
The optional
-ip(not shown in the example below) and-portoptions tell this H2O node what IP address and ports (port and port+1 are used) to use. The-ipoption is especially helpful for hosts that have multiple network interfaces.$ java -Xmx20g -jar h2o.jar -flatfile flatfile.txt -port 54321You will see output similar to the following:
05-11 16:40:46.268 172.16.2.39:54322 34242 main INFO: ----- H2O started ----- 05-11 16:40:46.337 172.16.2.39:54322 34242 main INFO: Build git branch: master 05-11 16:40:46.337 172.16.2.39:54322 34242 main INFO: Build git hash: 6c96387f893f3454912e20638dcb2f23a2786723 05-11 16:40:46.337 172.16.2.39:54322 34242 main INFO: Build git describe: jenkins-master-1192-10-g6c96387-dirty 05-11 16:40:46.337 172.16.2.39:54322 34242 main INFO: Build project version: 0.3.0.99999 05-11 16:40:46.337 172.16.2.39:54322 34242 main INFO: Built by: 'H2OUser' 05-11 16:40:46.337 172.16.2.39:54322 34242 main INFO: Built on: '2015-05-08 11:19:26' 05-11 16:40:46.337 172.16.2.39:54322 34242 main INFO: Java availableProcessors: 8 05-11 16:40:46.338 172.16.2.39:54322 34242 main INFO: Java heap totalMemory: 245.5 MB 05-11 16:40:46.338 172.16.2.39:54322 34242 main INFO: Java heap maxMemory: 17.78 GB 05-11 16:40:46.338 172.16.2.39:54322 34242 main INFO: Java version: Java 1.7.0_67 (from Oracle Corporation) 05-11 16:40:46.338 172.16.2.39:54322 34242 main INFO: OS version: Mac OS X 10.10.3 (x86_64) 05-11 16:40:46.338 172.16.2.39:54322 34242 main INFO: Machine physical memory: 16.00 GB 05-11 16:40:46.339 172.16.2.39:54322 34242 main INFO: X-h2o-cluster-id: 1431387646125 05-11 16:40:46.339 172.16.2.39:54322 34242 main INFO: Opted out of sending usage metrics. 05-11 16:40:46.339 172.16.2.39:54322 34242 main INFO: Possible IP Address: en5 (en5), fe80:0:0:0:daeb:97ff:feb3:6d4b%4 05-11 16:40:46.339 172.16.2.39:54322 34242 main INFO: Possible IP Address: en5 (en5), 172.16.2.39 05-11 16:40:46.339 172.16.2.39:54322 34242 main INFO: Possible IP Address: lo0 (lo0), fe80:0:0:0:0:0:0:1%1 05-11 16:40:46.339 172.16.2.39:54322 34242 main INFO: Possible IP Address: lo0 (lo0), 0:0:0:0:0:0:0:1 05-11 16:40:46.339 172.16.2.39:54322 34242 main INFO: Possible IP Address: lo0 (lo0), 127.0.0.1 05-11 16:40:46.340 172.16.2.39:54322 34242 main INFO: Internal communication uses port: 54323 05-11 16:40:46.340 172.16.2.39:54322 34242 main INFO: Listening for HTTP and REST traffic on http://172.16.2.39:54322/ 05-11 16:40:46.342 172.16.2.39:54322 34242 main INFO: H2O cloud name: 'H2OUser' on /172.16.2.39:54322, static configuration based on -flatfile flatfile.txt 05-11 16:40:46.342 172.16.2.39:54322 34242 main INFO: If you have trouble connecting, try SSH tunneling from your local machine (e.g., via port 55555): 05-11 16:40:46.342 172.16.2.39:54322 34242 main INFO: 1. Open a terminal and run 'ssh -L 55555:localhost:54322 H2OUser@172.16.2.39' 05-11 16:40:46.342 172.16.2.39:54322 34242 main INFO: 2. Point your browser to http://localhost:55555 05-11 16:40:46.542 172.16.2.39:54322 34242 main INFO: Log dir: '/tmp/h2o-H2OUser/h2ologs' 05-11 16:40:46.543 172.16.2.39:54322 34242 main INFO: Cur dir: '/Users/H2OUser/h2o-3' 05-11 16:40:46.564 172.16.2.39:54322 34242 main INFO: HDFS subsystem successfully initialized 05-11 16:40:46.565 172.16.2.39:54322 34242 main INFO: S3 subsystem successfully initialized 05-11 16:40:46.565 172.16.2.39:54322 34242 main INFO: Flow dir: '/Users/H2OUser/h2oflows' 05-11 16:40:46.578 172.16.2.39:54322 34242 main INFO: Cloud of size 3 formed [/172.16.2.39:54322, 172.16.2.40:54322, 172.16.2.41:54322]As you add more nodes to your cluster, the H2O output will inform you:
INFO: Cloud of size 3 formed [/...]Access the H2O Web UI with your browser. Point your browser to the IP address listed under “Listening for HTTP and REST traffic on…” in the H2O output.
If you are programmatically creating the cloud, give the cloud some time to establish itself (typically one minute is sufficient) and then check to see if the cloud is up.
To check the cloud’s status, point to the url http://
: /Cloud.json (see a piece of the JSON response below). Wait for cloud_sizeto be the expected value and theconsensusfield to be true.
{
...
"cloud_size": 2,
"consensus": true,
...
}
… On EC2 and S3
Note: If you would like to try out H2O on an EC2 cluster, play.h2o.ai is the easiest way to get started. H2O Play provides access to a temporary cluster managed by H2O.
If you would still like to set up your own EC2 cluster, follow the instructions below.
On EC2
Tested on Redhat AMI, Amazon Linux AMI, and Ubuntu AMI
To use the Amazon Web Services (AWS) S3 storage solution, you will need to pass your S3 access credentials to H2O. This will allow you to access your data on S3 when importing data frames with path prefixes s3n://....
For security reasons, we recommend writing a script to read the access credentials that are stored in a separate file. This will not only keep your credentials from propagating to other locations, but it will also make it easier to change the credential information later.
Standalone Instance
When running H2O in standalone mode using the simple Java launch command, we can pass in the S3 credentials in two ways.
You can pass in credentials in standalone mode the same way as accessing data from HDFS on Hadoop. Create a
core-site.xmlfile and pass it in with the flag-hdfs_config. For an examplecore-site.xmlfile, refer to Core-site.xml.Edit the properties in the core-site.xml file to include your Access Key ID and Access Key as shown in the following example:
<property> <name>fs.s3n.awsAccessKeyId</name> <value>[AWS SECRET KEY]</value> </property> <property> <name>fs.s3n.awsSecretAccessKey</name> <value>[AWS SECRET ACCESS KEY]</value> </property>Launch with the configuration file
core-site.xmlby entering the following in the command line:java -jar h2o.jar -hdfs_config core-site.xml
Import the data using importFile with the S3 url path:
s3n://bucket/path/to/file.csv
You can pass the AWS Access Key and Secret Access Key in an S3N Url in Flow, R, or Python (where
AWS_ACCESS_KEYrepresents your user name andAWS_SECRET_KEYrepresents your password).To import the data from the Flow API:
`importFiles [ "s3n://<AWS_ACCESS_KEY>:<AWS_SECRET_KEY>@bucket/path/to/file.csv" ]`To import the data from the R API:
`h2o.importFile(path = "s3n://<AWS_ACCESS_KEY>:<AWS_SECRET_KEY>@bucket/path/to/file.csv")`To import the data from the Python API:
`h2o.import_frame(path = "s3n://<AWS_ACCESS_KEY>:<AWS_SECRET_KEY>@bucket/path/to/file.csv")`
Core-site.xml Example
The following is an example core-site.xml file:
<?xml version="1.0"?>
<?xml-stylesheet type="text/xsl" href="configuration.xsl"?>
<!-- Put site-specific property overrides in this file. -->
<configuration>
<!--
<property>
<name>fs.default.name</name>
<value>s3n://<your s3 bucket></value>
</property>
-->
<property>
<name>fs.s3n.awsAccessKeyId</name>
<value>insert access key here</value>
</property>
<property>
<name>fs.s3n.awsSecretAccessKey</name>
<value>insert secret key here</value>
</property>
</configuration>
Launching H2O
- Selecting the Operating System and Virtualization Type
- Configuring the Instance
- Downloading Java and H2O
Note: Before launching H2O on an EC2 cluster, verify that ports 54321 and 54322 are both accessible by TCP and UDP.
Selecting the Operating System and Virtualization Type
Select your operating system and the virtualization type of the prebuilt AMI on Amazon. If you are using Windows, you will need to use a hardware-assisted virtual machine (HVM). If you are using Linux, you can choose between para-virtualization (PV) and HVM. These selections determine the type of instances you can launch.
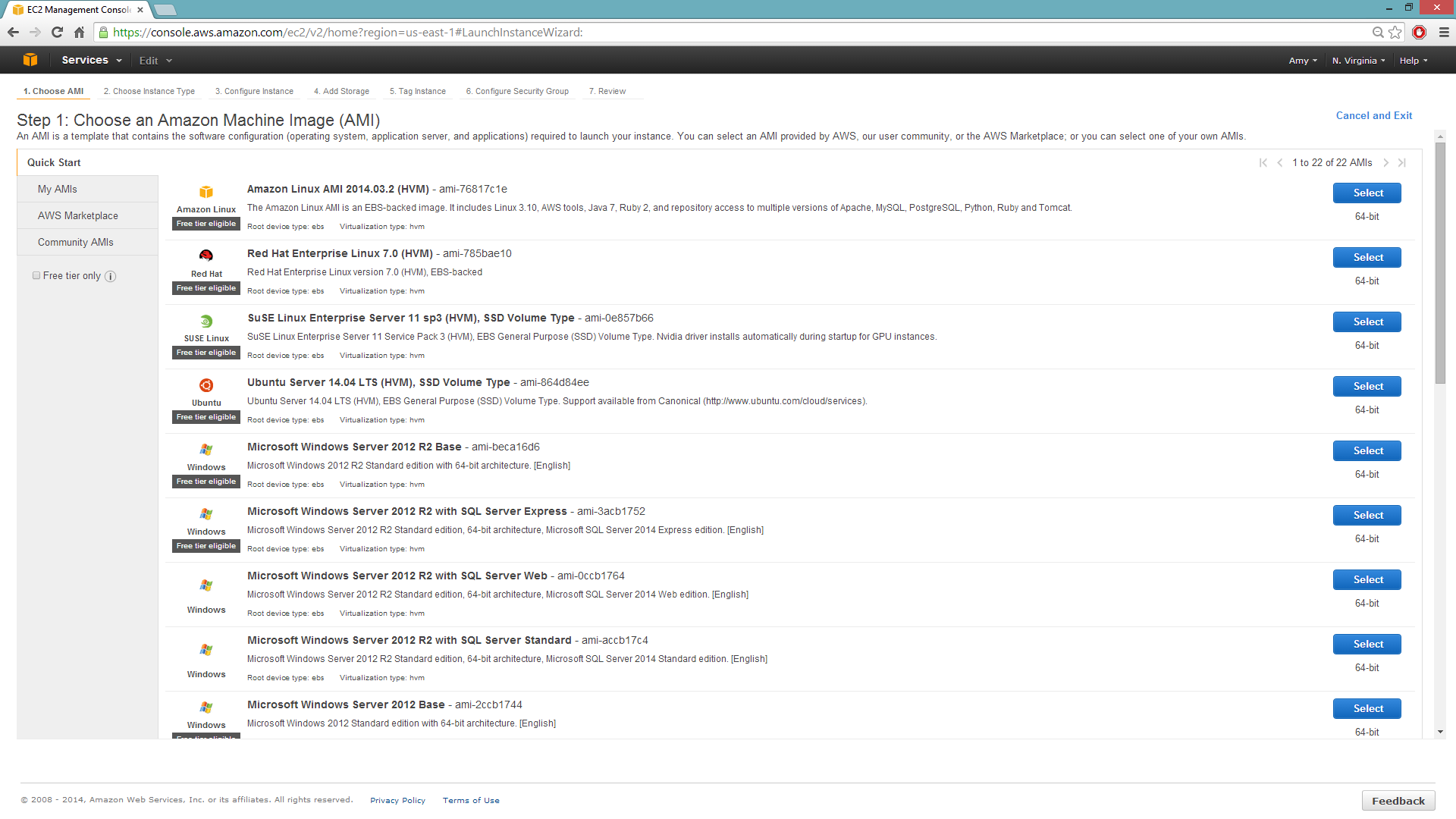
For more information about virtualization types, refer to Amazon.
Configuring the Instance
Select the IAM role and policy to use to launch the instance. H2O detects the temporary access keys associated with the instance, so you don’t need to copy your AWS credentials to the instances.
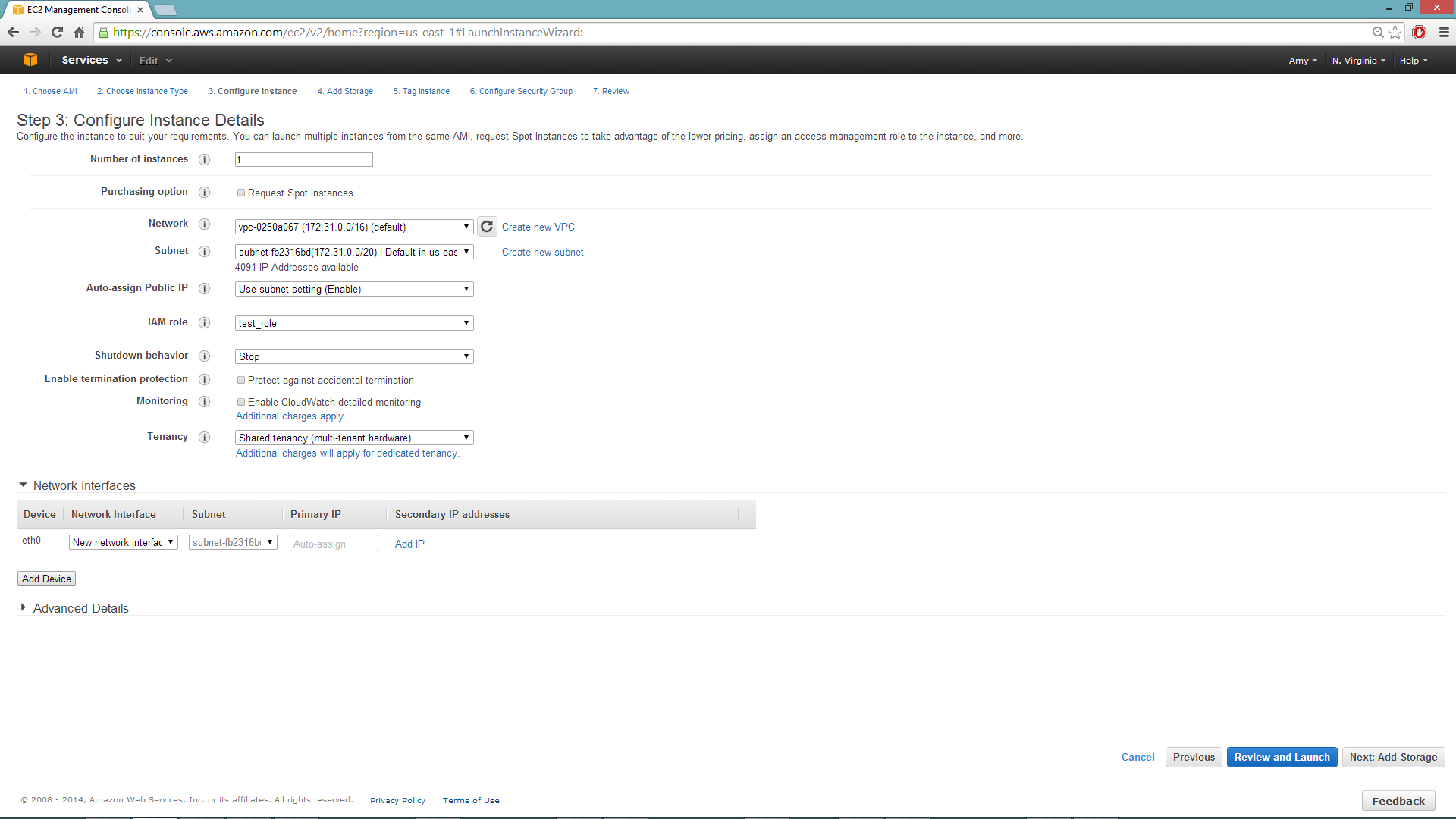
When launching the instance, select an accessible key pair.
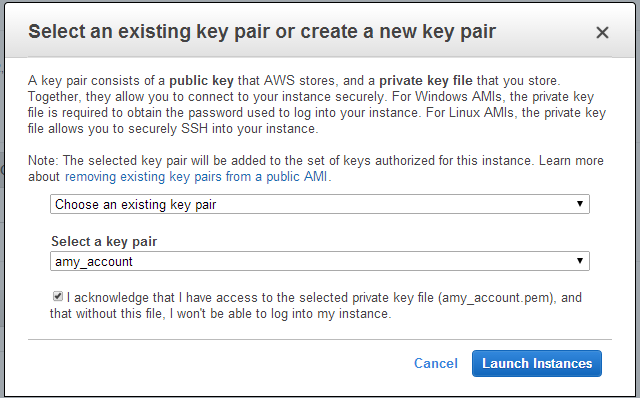
(Windows Users) Tunneling into the Instance
For Windows users that do not have the ability to use ssh from the terminal, either download Cygwin or a Git Bash that has the capability to run ssh:
ssh -i amy_account.pem ec2-user@54.165.25.98
Otherwise, download PuTTY and follow these instructions:
- Launch the PuTTY Key Generator.
- Load your downloaded AWS pem key file. Note: To see the file, change the browser file type to “All”.
Save the private key as a .ppk file.
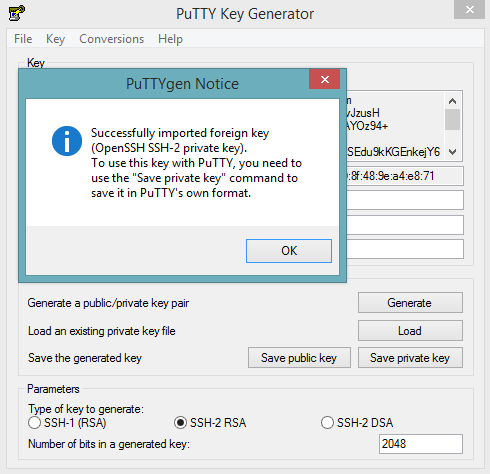
Launch the PuTTY client.
In the Session section, enter the host name or IP address. For Ubuntu users, the default host name is
ubuntu@<ip-address>. For Linux users, the default host name isec2-user@<ip-address>.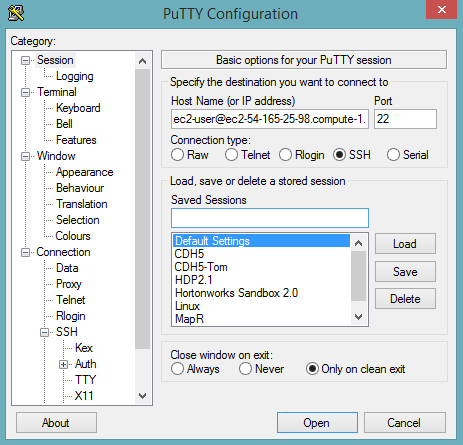
Select SSH, then Auth in the sidebar, and click the Browse button to select the private key file for authentication.
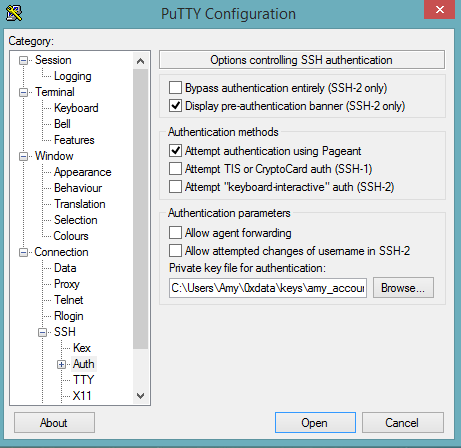
Start a new session and click the Yes button to confirm caching of the server’s rsa2 key fingerprint and continue connecting.
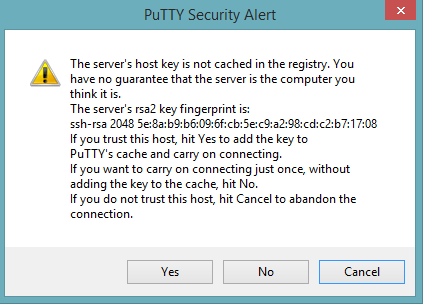
Downloading Java and H2O
- Download Java (JDK 1.7 or later) if it is not already available on the instance.
To download H2O, run the
wgetcommand with the link to the zip file available on our website by copying the link associated with the Download button for the selected H2O build.wget http://h2o-release.s3.amazonaws.com/h2o/rel-shannon/1/index.html unzip h2o-0.2.1.1.zip cd h2o-0.2.1.1 java -Xmx4g -jar h2o.jar- From your browser, navigate to
<Private_IP_Address>:54321or<Public_DNS>:54321to use H2O’s web interface.
… On Hadoop
Currently supported versions:
- CDH 5.2
- CDH 5.3
- HDP 2.1
- HDP 2.2
- MapR 3.1.1
- MapR 4.0.1
Important Points to Remember:
- Each H2O node runs as a mapper
- Run only one mapper per host
- There are no combiners or reducers
- Each H2O cluster must have a unique job name
-mapperXmx,-nodes, and-outputare required- Root permissions are not required - just unzip the H2O .zip file on any single node
Prerequisite: Open Communication Paths
H2O communicates using two communication paths. Verify these are open and available for use by H2O.
Path 1: mapper to driver
Optionally specify this port using the -driverport option in the hadoop jar command (see “Hadoop Launch Parameters” below). This port is opened on the driver host (the host where you entered the hadoop jar command). By default, this port is chosen randomly by the operating system.
Path 2: mapper to mapper
Optionally specify this port using the -baseport option in the hadoop jar command (see “Hadoop Launch Parameters” below). This port and the next subsequent port are opened on the mapper hosts (the Hadoop worker nodes) where the H2O mapper nodes are placed by the Resource Manager. By default, ports 54321 (TCP) and 54322 (TCP & UDP) are used.
The mapper port is adaptive: if 54321 and 54322 are not available, H2O will try 54323 and 54324 and so on. The mapper port is designed to be adaptive because sometimes if the YARN cluster is low on resources, YARN will place two H2O mappers for the same H2O cluster request on the same physical host. For this reason, we recommend opening a range of more than two ports (20 ports should be sufficient).
Tutorial
The following tutorial will walk the user through the download or build of H2O and the parameters involved in launching H2O from the command line.
Download the latest H2O release for your version of Hadoop:
wget http://h2o-release.s3.amazonaws.com/h2o/master/1110/h2o-0.3.0.1110-cdh5.2.zip wget http://h2o-release.s3.amazonaws.com/h2o/master/1110/h2o-0.3.0.1110-cdh5.3.zip wget http://h2o-release.s3.amazonaws.com/h2o/master/1110/h2o-0.3.0.1110-hdp2.1.zip wget http://h2o-release.s3.amazonaws.com/h2o/master/1110/h2o-0.3.0.1110-hdp2.2.zip wget http://h2o-release.s3.amazonaws.com/h2o/master/1110/h2o-0.3.0.1110-mapr3.1.1.zip wget http://h2o-release.s3.amazonaws.com/h2o/master/1110/h2o-0.3.0.1110-mapr4.0.1.zipNote: Enter only one of the above commands.
Prepare the job input on the Hadoop Node by unzipping the build file and changing to the directory with the Hadoop and H2O’s driver jar files.
unzip h2o-0.3.0.1110-*.zip cd h2o-0.3.0.1110-*To launch H2O nodes and form a cluster on the Hadoop cluster, run:
hadoop jar h2odriver.jar -nodes 1 -mapperXmx 1g -output hdfsOutputDirNameThe above command launches a 1g node of H2O. We recommend you launch the cluster with at least four times the memory of your data file size.
mapperXmx is the mapper size or the amount of memory allocated to each node.
nodes is the number of nodes requested to form the cluster.
output is the name of the directory created each time a H2O cloud is created so it is necessary for the name to be unique each time it is launched.
To monitor your job, direct your web browser to your standard job tracker Web UI. To access H2O’s Web UI, direct your web browser to one of the launched instances. If you are unsure where your JVM is launched, review the output from your command after the nodes has clouded up and formed a cluster. Any of the nodes’ IP addresses will work as there is no master node.
Determining driver host interface for mapper->driver callback... [Possible callback IP address: 172.16.2.181] [Possible callback IP address: 127.0.0.1] ... Waiting for H2O cluster to come up... H2O node 172.16.2.184:54321 requested flatfile Sending flatfiles to nodes... [Sending flatfile to node 172.16.2.184:54321] H2O node 172.16.2.184:54321 reports H2O cluster size 1 H2O cluster (1 nodes) is up Blocking until the H2O cluster shuts down...
Hadoop Launch Parameters
-h | -help: Display help-jobname <JobName>: Specify a job name for the Jobtracker to use; the default isH2O_nnnnn(where n is chosen randomly)-driverif <IP address of mapper -> driver callback interface>: Specify the IP address for callback messages from the mapper to the driver.-driverport <port of mapper -> callback interface>: Specify the port number for callback messages from the mapper to the driver.-network <IPv4Network1>[,<IPv4Network2>]: Specify the IPv4 network(s) to bind to the H2O nodes; multiple networks can be specified to force H2O to use the specified host in the Hadoop cluster.10.1.2.0/24allows 256 possibilities.-timeout <seconds>: Specify the timeout duration (in seconds) to wait for the cluster to form before failing.-disown: Exit the driver after the cluster forms.notify <notification file name>: Specify a file to write when the cluster is up. The file contains the IP and port of the embedded web server for one of the nodes in the cluster. All mappers must start before the H2O cloud is considered “up”.mapperXmx <per mapper Java Xmx heap size>: Specify the amount of memory to allocate to H2O.extramempercent <0-20>: Specify the extra memory for internal JVM use outside of the Java heap. This is a percentage ofmapperXmx.-n | -nodes <number of H2O nodes>: Specify the number of nodes.-nthreads <maximum number of CPUs>: Specify the number of CPUs to use. Enter-1to use all CPUs on the host, or enter a positive integer.-baseport <initialization port for H2O nodes>: Specify the initialization port for the H2O nodes. The default is54321.-ea: Enable assertions to verify boolean expressions for error detection.-verbose:gc: Include heap and garbage collection information in the logs.-XX:+PrintGCDetails: Include a short message after each garbage collection.-license <license file name>: Specify the directory of local filesytem location and the license file name.-o | -output <HDFS output directory>: Specify the HDFS directory for the output.-flow_dir <Saved Flows directory>: Specify the directory for saved flows. By default, H2O will try to find the HDFS home directory to use as the directory for flows. If the HDFS home directory is not found, flows cannot be saved unless a directory is specified using-flow_dir.
Accessing S3 Data from Hadoop
H2O launched on Hadoop can access S3 Data in addition to to HDFS. To enable access, follow the instructions below.
Edit Hadoop’s core-site.xml, then set the HADOOP_CONF_DIR environment property to the directory containing the core-site.xml file. For an example core-site.xml file, refer to Core-site.xml. Typically, the configuration directory for most Hadoop distributions is /etc/hadoop/conf.
You can also pass the S3 credentials when launching H2O with the Hadoop jar command. Use the -D flag to pass the credentials:
hadoop jar h2odriver.jar -Dfs.s3.awsAccessKeyId="${AWS_ACCESS_KEY}" -Dfs.s3n.awsSecretAccessKey="${AWS_SECRET_KEY}" -n 3 -mapperXmx 10g -output outputDirectory
where AWS_ACCESS_KEY represents your user name and AWS_SECRET_KEY represents your password.
Then import the data with the S3 URL path:
To import the data from the Flow API:
importFiles [ "s3n://bucket/path/to/file.csv" ]To import the data from the R API:
h2o.importFile(path = "s3n://bucket/path/to/file.csv")To import the data from the Python API:
h2o.import_frame(path = "s3n://bucket/path/to/file.csv")
… Using Docker
This walkthrough describes:
- Installing Docker on Mac or Linux OS
- Creating and modifying the Dockerfile
- Building a Docker image from the Dockerfile
- Running the Docker build
- Launching H2O
- Accessing H2O from the web browser or R
Walkthrough
Prerequisites
Linux kernel version 3.8+
or
Mac OS X 10.6+
Note: Older Linux kernel versions are known to cause kernel panics and to break Docker; there are ways around it, but attempt at your own risk.
You can check the version of your kernel by running uname -r in your terminal. The following walkthrough has been tested on a Mac OS X 10.10.1.
Step 1 - Install and Launch Docker
Step 2 - Create or Download Dockerfile
Create a folder on the Host OS to host your Dockerfile by running:
mkdir -p /data/h2o-shannon
Then either download or create a Dockerfile. The Dockerfile is essentially a build recipe that will be used to build the container.
Download and use our Dockerfile template by running:
cd /data/h2o-shannon
wget http://h2o.ai/blog/2015_01_h2o-docker/Dockerfile
The Dockerfile will:
- Pull and update the base image (Ubuntu 14.04)
- Install Java 7
- Fetch and download the H2O Shannon build from H2O’s S3 repository
- Expose port 54321 and 54322 in preparation for launching H2O on those ports
Step 3 - Build Docker image from Dockerfile
From the /data/h2o-shannon directory, run:
docker build -t="h2o.ai/shannon" .
This process can take a few minutes as it assembles all the necessary parts to the image.
Step 4 - Run Docker Build
On a Mac, you must use the argument -p 54321:54321 to expressly map the port 54321. This is redundant on Linux.
docker run -it -p 54321:54321 h2o.ai/shannon
Step 5 - Launch H2O
Step into the /opt directory and launch H2O. Change the value of -Xmx to the amount of memory you want to allocate to the H2O instance. By default, H2O launches on port 54321.
cd /opt
java -Xmx1g -jar h2o.jar
Step 6 - Access H2O from the web browser or R
- On Linux, when H2O finishes launching, you can copy and paste the IP address and port of the H2O instance. In the following example, that would be 172.17.0.5:54321.
03:58:25.963 main INFO WATER: Cloud of size 1 formed [/172.17.0.5:54321 (00:00:00.000)]
- If it is running on a Mac, you will need to find the IP address of the Docker’s network that bridges to your Host OS. To do this, open a new terminal (not a bash for your container) and run
boot2docker ip.
$ boot2docker ip
192.168.59.103
Once you have the IP address, point your browser to the specified ip address and port. In R, you can access the instance by installing the latest version of the H2O R package and running:
library(h2o)
dockerH2O <- h2o.init(ip = "192.168.59.103", port = 54321)
Flow Web UI …
H2O Flow is an open-source user interface for H2O. It is a web-based interactive environment that allows you to combine code execution, text, mathematics, plots, and rich media in a single document, similar to iPython Notebooks.
With H2O Flow, you can capture, rerun, annotate, present, and share your workflow. H2O Flow allows you to use H2O interactively to import files, build models, and iteratively improve them. Based on your models, you can make predictions and add rich text to create vignettes of your work - all within Flow’s browser-based environment.
Flow’s hybrid user interface seamlessly blends command-line computing with a modern graphical user interface. However, rather than displaying output as plain text, Flow provides a point-and-click user interface for every H2O operation. It allows you to access any H2O object in the form of well-organized tabular data.
H2O Flow sends commands to H2O as a sequence of executable cells. The cells can be modified, rearranged, or saved to a library. Each cell contains an input field that allows you to enter commands, define functions, call other functions, and access other cells or objects on the page. When you execute the cell, the output is a graphical object, which can be inspected to view additional details.
While H2O Flow supports REST API, R scripts, and CoffeeScript, no programming experience is required to run H2O Flow. You can click your way through any H2O operation without ever writing a single line of code. You can even disable the input cells to run H2O Flow using only the GUI. H2O Flow is designed to guide you every step of the way, by providing input prompts, interactive help, and example flows.
Introduction
This guide will walk you through how to use H2O’s web UI, H2O Flow. To view a demo video of H2O Flow, click here.
Getting Help
First, let’s go over the basics. Type h to view a list of helpful shortcuts.
The following help window displays:

To close this window, click the X in the upper-right corner, or click the Close button in the lower-right corner. You can also click behind the window to close it. You can also access this list of shortcuts by clicking the Help menu and selecting Keyboard Shortcuts.
For additional help, select the Help sidebar to the right and click the Assist Me! button.
You can also type assist in a blank cell and press Ctrl+Enter. A list of common tasks displays to help you find the correct command.
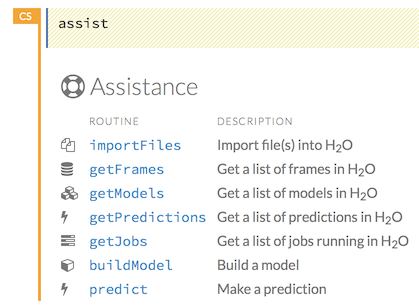
There are multiple resources to help you get started with Flow in the Help sidebar. To access this document, select the Getting Started with H2O Flow link below the Help Topics heading.
You can also explore the pre-configured flows available in H2O Flow for a demonstration of how to create a flow. To view the example flows, click the Browse installed packs… link in the Packs subsection of the Help sidebar. Click the examples folder and select the example flow from the list.
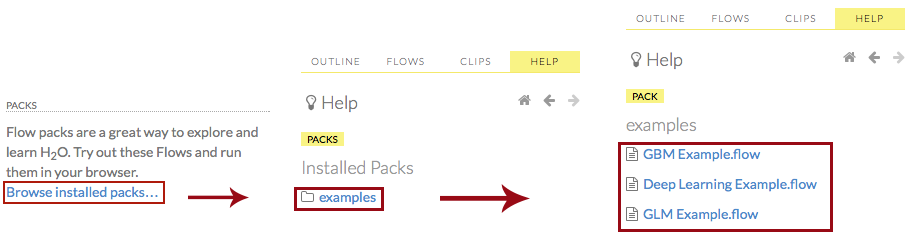
If you have a flow currently open, a confirmation window appears asking if the current notebook should be replaced. To load the example flow, click the Load Notebook button.
To view the REST API documentation, click the Help tab in the sidebar and then select the type of REST API documentation (Routes or Schemas).

Before getting started with H2O Flow, make sure you understand the different cell modes.
Understanding Cell Modes
There are two modes for cells: edit and command.
Using Edit Mode
In edit mode, the cell is yellow with a blinking bar to indicate where text can be entered and there is an orange flag to the left of the cell.

Using Command Mode
In command mode, the flag is yellow. The flag also indicates the cell’s format:
MD: Markdown
Note: Markdown formatting is not applied until you run the cell by clicking the Run button or clicking the Run menu and selecting Run.

CS: Code (default)
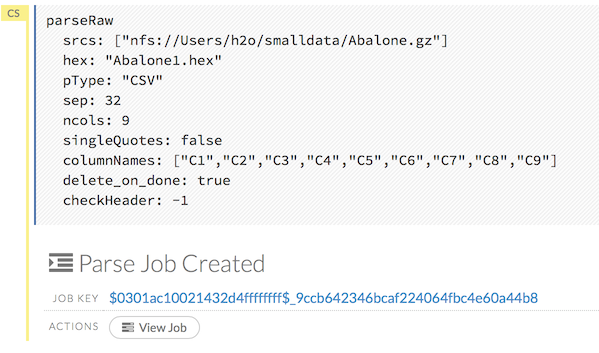
RAW: Raw format (for code comments)

H[1-6]: Heading level (where 1 is a first-level heading)

NOTE: If there is an error in the cell, the flag is red.
If the cell is executing commands, the flag is teal. The flag returns to yellow when the task is complete.
Changing Cell Formats
To change the cell’s format (for example, from code to Markdown), make sure you are in not in command (not edit) mode and that the cell you want to change is selected. The easiest way to do this is to click on the flag to the left of the cell. Enter the keyboard shortcut for the format you want to use. The flag’s text changes to display the current format.
| Cell Mode | Keyboard Shortcut |
|---|---|
| Code | y |
| Markdown | m |
| Raw text | r |
| Heading 1 | 1 |
| Heading 2 | 2 |
| Heading 3 | 3 |
| Heading 4 | 4 |
| Heading 5 | 5 |
| Heading 6 | 6 |
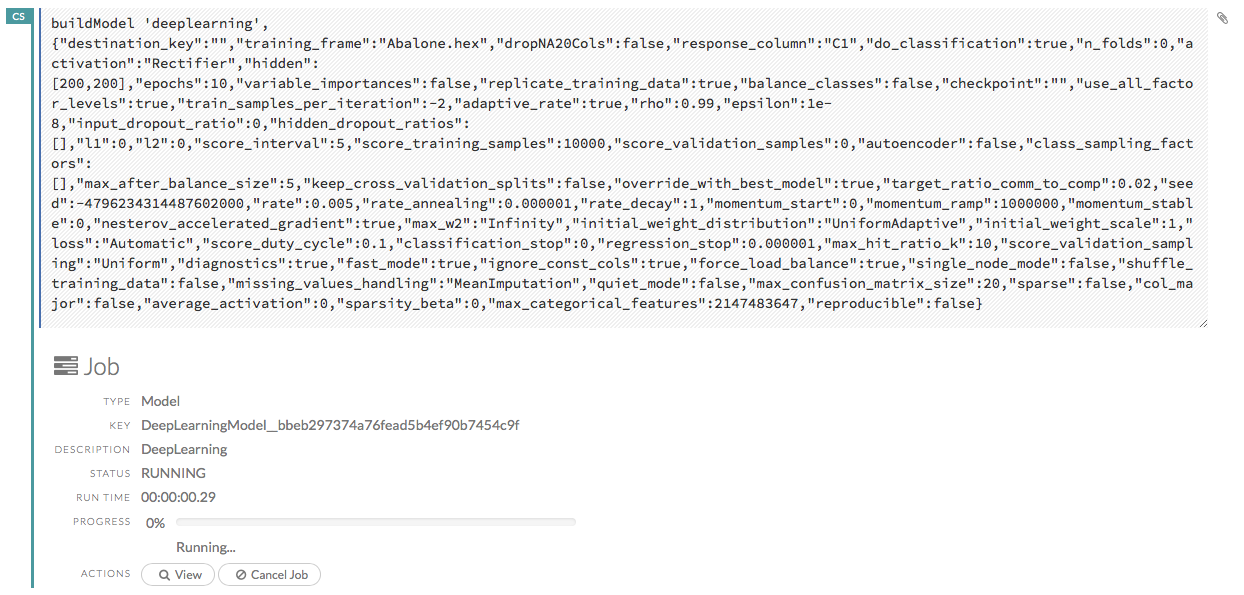
Running Flows
When you run the flow, a progress bar that indicates the current status of the flow. You can cancel the currently running flow by clicking the Stop button in the progress bar.
When the flow is complete, a message displays in the upper right. Note: If there is an error in the flow, H2O Flow stops the flow at the cell that contains the error.
Using Keyboard Shortcuts
Here are some important keyboard shortcuts to remember:
- Click a cell and press Enter to enter edit mode, which allows you to change the contents of a cell.
- To exit edit mode, press Esc.
- To execute the contents of a cell, press the Ctrl and Enter buttons at the same time.
The following commands must be entered in command mode.
- To add a new cell above the current cell, press a.
- To add a new cell below the current cell, press b.
- To delete the current cell, press the d key twice. (dd).
You can view these shortcuts by clicking Help > Keyboard Shortcuts or by clicking the Help tab in the sidebar.
Using Flow Buttons
There are also a series of buttons at the top of the page below the flow name that allow you to save the current flow, add a new cell, move cells up or down, run the current cell, and cut, copy, or paste the current cell. If you hover over the button, a description of the button’s function displays.

You can also use the menus at the top of the screen to edit the order of the cells, toggle specific format types (such as input or output), create models, or score models. You can also access troubleshooting information or obtain help with Flow.

Note: To disable the code input and use H2O Flow strictly as a GUI, click the Cell menu, then Toggle Cell Input.
Now that you are familiar with the cell modes, let’s import some data.
… Importing Data
If you don’t have any of your own data to work with, you can find some example datasets here:
There are multiple ways to import data in H2O flow:
Click the Assist Me! button in the Help sidebar, then click the importFiles link. Enter the file path in the auto-completing Search entry field and press Enter. Select the file from the search results and select it by clicking the Add All link.
You can also drag and drop the file onto the Search field in the cell.
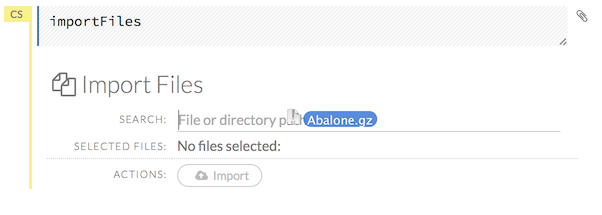
In a blank cell, select the CS format, then enter
importFiles ["path/filename.format"](wherepath/filename.formatrepresents the complete file path to the file, including the full file name. The file path can be a local file path or a website address.
After selecting the file to import, the file path displays in the “Search Results” section. To import a single file, click the plus sign next to the file. To import all files in the search results, click the Add all link. The files selected for import display in the “Selected Files” section.
Note: If the file is compressed, it will only be read using a single thread. For best performance, we recommend uncompressing the file before importing, as this will allow use of the faster multithreaded distributed parallel reader during import.
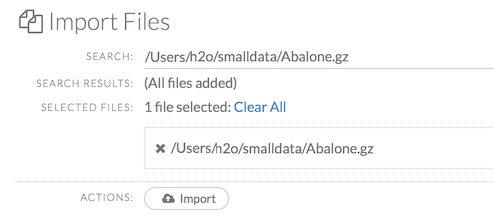
To import the selected file(s), click the Import button.
To remove all files from the “Selected Files” list, click the Clear All link.
To remove a specific file, click the X next to the file path.
After you click the Import button, the raw code for the current job displays. A summary displays the results of the file import, including the number of imported files and their Network File System (nfs) locations.
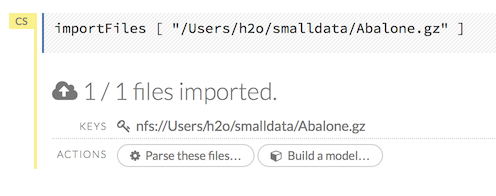
Uploading Data
To upload a local file, click the Data menu and select Upload File…. Click the Choose File button, select the file, click the Choose button, then click the Upload button.
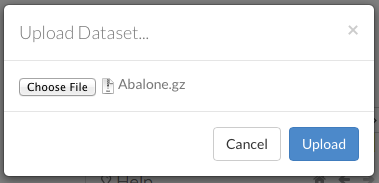
When the file has uploaded successfully, a message displays in the upper right and the Setup Parse cell displays.

Ok, now that your data is available in H2O Flow, let’s move on to the next step: parsing. Click the Parse these files button to continue.
Parsing Data
After you have imported your data, parse the data.
Select the parser type (if necessary) from the drop-down Parser list. For most data parsing, H2O automatically recognizes the data type, so the default settings typically do not need to be changed. The following options are available:
- Auto
- ARFF
- XLS
- XLSX
- CSV
- SVMLight
If a separator or delimiter is used, select it from the Separator list.
Select a column header option, if applicable:
- Auto: Automatically detect header types.
- First row contains column names: Specify heading as column names.
- First row contains data: Specify heading as data. This option is selected by default.
Select any necessary additional options:
- Enable single quotes as a field quotation character: Treat single quote marks (also known as apostrophes) in the data as a character, rather than an enum. This option is not selected by default.
- Delete on done: Check this checkbox to delete the imported data after parsing. This option is selected by default.
A preview of the data displays in the “Data Preview” section.
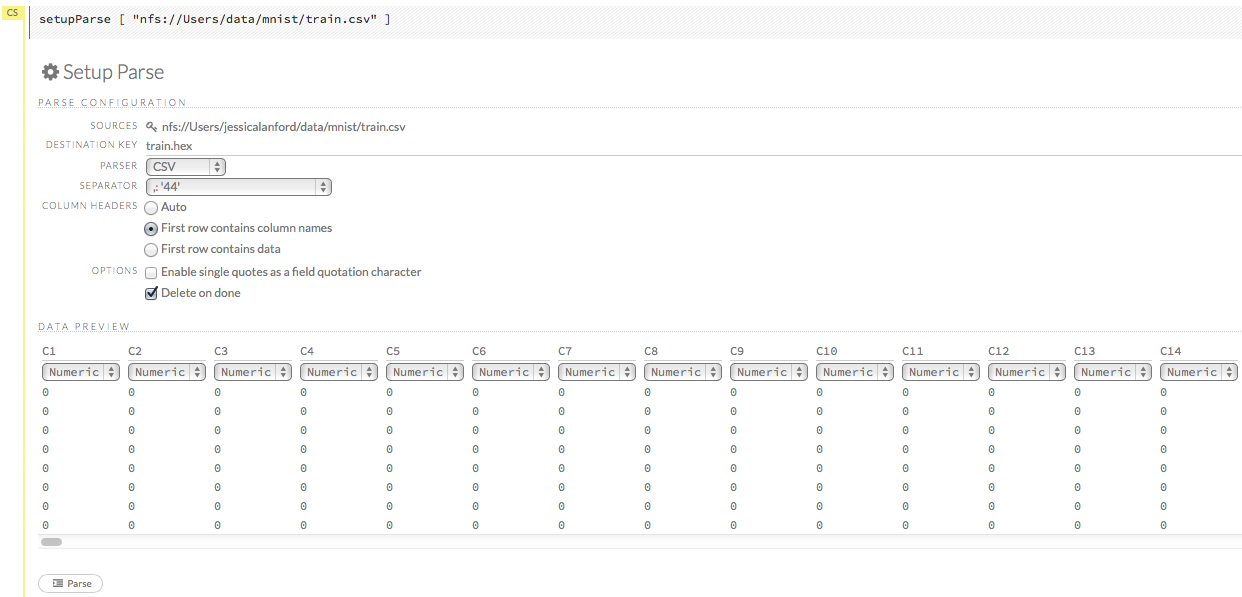
Note: To change the column type, select the drop-down list at the top of the column and select the data type. The options are:
- Unknown
- Numeric
- Enum
- Time
- UUID
- String
- Invalid
After making your selections, click the Parse button.
After you click the Parse button, the code for the current job displays.

Since we’ve submitted a couple of jobs (data import & parse) to H2O now, let’s take a moment to learn more about jobs in H2O.
Viewing Jobs
Any command (such as importFiles) you enter in H2O is submitted as a job, which is associated with a key. The key identifies the job within H2O and is used as a reference.
Viewing All Jobs
To view all jobs, click the Admin menu, then click Jobs, or enter getJobs in a cell in CS mode.
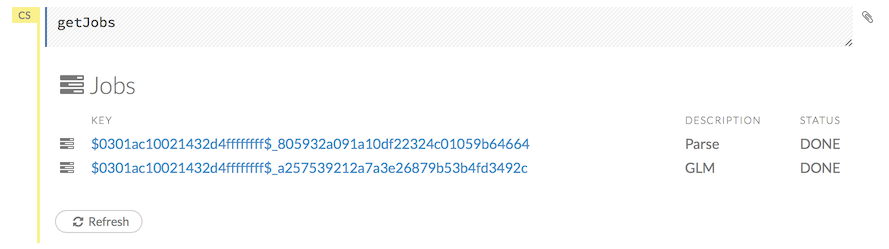
The following information displays:
- Type (for example,
FrameorModel) - Link to the object
- Description of the job type (for example,
ParseorGBM) - Start time
- End time
- Run time
To refresh this information, click the Refresh button. To view the details of the job, click the View button.
Viewing Specific Jobs
To view a specific job, click the link in the “Destination” column.
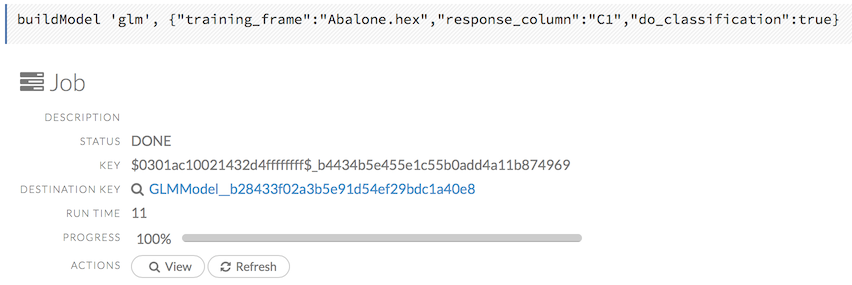
The following information displays:
- Type (for example,
Frame) - Link to object (key)
- Description (for example,
Parse) - Status
- Run time
- Progress
NOTE: For a better understanding of how jobs work, make sure to review the Viewing Frames section as well.
Ok, now that you understand how to find jobs in H2O, let’s submit a new one by building a model.
… Building Models
To build a model:
Click the Assist Me! button and select buildModel
or
Click the Assist Me! button, select getFrames, then click the Build Model… button below the parsed .hex data set
or
Click the View button after parsing data, then click the Build Model button
or
Click the drop-down Model menu and select the model type from the list
The Build Model… button can be accessed from any page containing the .hex key for the parsed data (for example, getJobs > getFrame).
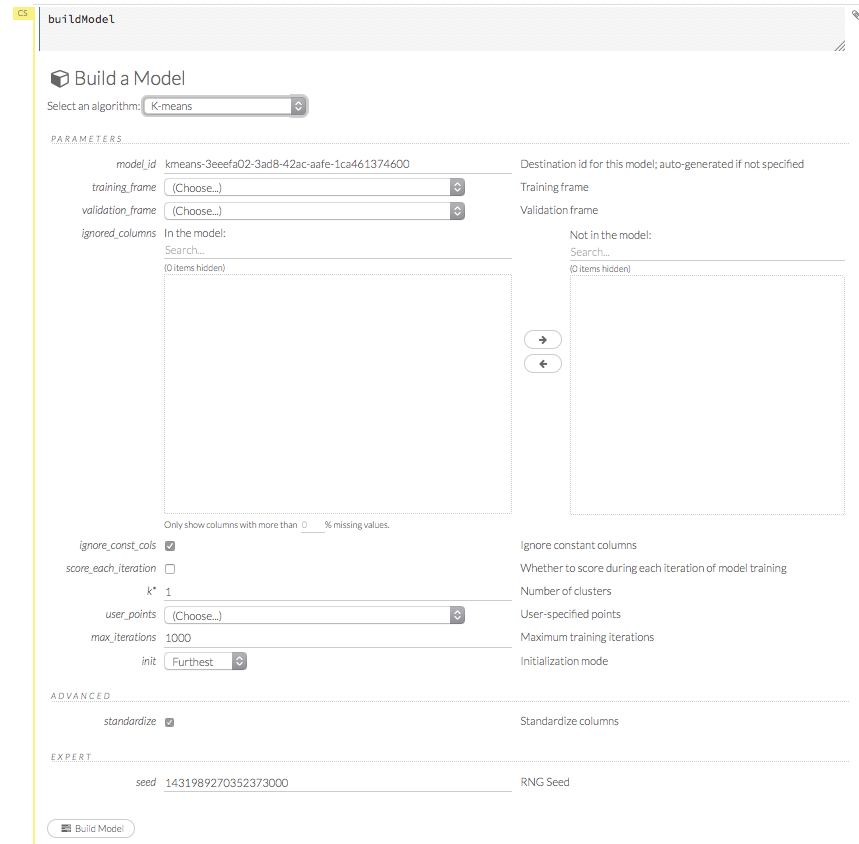
In the Build a Model cell, select an algorithm from the drop-down menu:
- K-means: Create a K-Means model.
- Generalized Linear Model: Create a Generalized Linear model.
- Distributed RF: Create a distributed Random Forest model.
- Naïve Bayes: Create a Naïve Bayes model.
- Principal Component Analysis: Create a Principal Components Analysis model for modeling without regularization or performing dimensionality reduction.
- Gradient Boosting Machine: Create a Gradient Boosted model
- Deep Learning: Create a Deep Learning model.
The available options vary depending on the selected model. If an option is only available for a specific model type, the model type is listed. If no model type is specified, the option is applicable to all model types.
Model_ID: (Optional) Enter a custom name for the model to use as a reference. By default, H2O automatically generates an ID containing the model type (for example,
gbm-6f6bdc8b-ccbc-474a-b590-4579eea44596).Training_frame: (Required) Select the dataset used to build the model.
NOTE: If you click the Build a model button from the
Parsecell, the training frame is entered automatically.Validation_frame: (Optional) Select the dataset used to evaluate the accuracy of the model.
Ignored_columns: (Optional) Click the plus sign next to a column name to add it to the list of columns excluded from the model. To add all columns, click the -> button. To remove a column from the list of ignored columns, click the X next to the column name. To remove all columns from the list of ignored columns, click the <- button. To search for a specific column, type the column name in the Search field above the column list. To only show columns with a specific percentage of missing values, specify the percentage in the Only show columns with more than 0% missing values field.
User_points: (K-Means, PCA) For K-Means, specify the number of initial cluster centers. For PCA, specify the initial Y matrix. Note: The PCA User_points parameter should only be used by advanced users for testing purposes.
Transform: (PCA) Select the transformation method for the training data: None, Standardize, Normalize, Demean, or Descale. The default is None.
Response_column: (Required for GLM, GBM, DL, DRF, Naïve Bayes) Select the column to use as the independent variable.
Solver: (GLM) Select the solver to use (IRLSM, L_BFGS, or auto). IRLSM is fast on on problems with small number of predictors and for lambda-search with L1 penalty, while L_BFGS scales better for datasets with many columns. The default is IRLSM.
Ntrees: (GBM, DRF) Specify the number of trees. The default value is 50.
Max_depth: (GBM, DRF) Specify the maximum tree depth. For GBM, the default value is 5. For DRF, the default value is 20.
Min_rows: (GBM), (DRF) Specify the minimum number of observations for a leaf (“nodesize” in R). For Grid Search, use comma-separated values. The default value is 10.
Nbins: (GBM, DRF) Specify the number of bins for the histogram. The default value is 20.
Mtries: (DRF) Specify the columns to randomly select at each level. To use the square root of the columns, enter
-1. The default value is -1.Sample_rate: (DRF) Specify the sample rate. The range is 0 to 1.0 and the default value is 0.632.
Build_tree_one_node: (DRF) To run on a single node, check this checkbox. This is suitable for small datasets as there is no network overhead but fewer CPUs are used. The default setting is disabled.
Learn_rate: (GBM) Specify the learning rate. The range is 0.0 to 1.0 and the default is 0.1.
Distribution: (GBM) Select the distribution type from the drop-down list. The options are auto, bernoulli, multinomial, or gaussian and the default is auto.
Loss: (DL) Select the loss function. For DL, the options are Automatic, MeanSquare, CrossEntropy, Huber, or Absolute and the default value is Automatic. Absolute, MeanSquare, and Huber are applicable for regression or classification, while CrossEntropy is only applicable for classification. Huber can improve for regression problems with outliers.
Score_each_iteration: (K-Means, DRF, Naïve Bayes, PCA, GBM) To score during each iteration of the model training, check this checkbox.
K: (K-Means), (PCA) For K-Means, specify the number of clusters. For PCA, specify the rank of matrix approximation. The default for K-Means and PCA is 1.
Gamma: (PCA) Specify the regularization weight for PCA. The default is 0.
Max_iterations: (K-Means, PCA,GLM) Specify the number of training iterations. For K-Means and PCA, the default is 1000. For GLM, the default is -1.
Beta_epsilon: (GLM) Specify the beta epsilon value. If the L1 normalization of the current beta change is below this threshold, consider using convergence.
Init: (K-Means, PCA) Select the initialization mode. For K-Means, the options are Furthest, PlusPlus, Random, or User. For PCA, the options are PlusPlus, User, or None.
Note: If PlusPlus is selected, the initial Y matrix is chosen by the final cluster centers from the K-Means PlusPlus algorithm.
Family: (GLM) Select the model type (Gaussian, Binomial, Poisson, or Gamma).
Activation: (DL) Select the activation function (Tanh, TanhWithDropout, Rectifier, RectifierWithDropout, Maxout, MaxoutWithDropout). The default option is Rectifier.
Hidden: (DL) Specify the hidden layer sizes (e.g., 100,100). For Grid Search, use comma-separated values: (10,10),(20,20,20). The default value is [200,200]. The specified value(s) must be positive.
Epochs: (DL) Specify the number of times to iterate (stream) the dataset. The value can be a fraction. The default value for DL is 10.0.
Variable_importances: (DL) Check this checkbox to compute variable importance. This option is not selected by default.
Laplace: (Naïve Bayes) Specify the Laplace smoothing parameter. The default value is 0.
Min_sdev: (Naïve Bayes) Specify the minimum standard deviation to use for observations without enough data. The default value is 0.001.
Eps_sdev: (Naïve Bayes) Specify the threshold for standard deviation. If this threshold is not met, the min_sdev value is used. The default value is 0.
Min_prob: (Naïve Bayes) Specify the minimum probability to use for observations without enough data. The default value is 0.001.
Eps_prob: (Naïve Bayes) Specify the threshold for standard deviation. If this threshold is not met, the min_sdev value is used. The default value is 0.
Standardize: (K-Means, GLM) To standardize the numeric columns to have mean of zero and unit variance, check this checkbox. Standardization is highly recommended; if you do not use standardization, the results can include components that are dominated by variables that appear to have larger variances relative to other attributes as a matter of scale, rather than true contribution. This option is selected by default.
Beta_constraints: (GLM)To use beta constraints, select a dataset from the drop-down menu. The selected frame is used to constraint the coefficient vector to provide upper and lower bounds.
Advanced Options
Checkpoint: (DL) Enter a model key associated with a previously-trained Deep Learning model. Use this option to build a new model as a continuation of a previously-generated model (e.g., by a grid search).
Use_all_factor_levels: (DL) Check this checkbox to use all factor levels in the possible set of predictors; if you enable this option, sufficient regularization is required. By default, the first factor level is skipped. For Deep Learning models, this option is useful for determining variable importances and is automatically enabled if the autoencoder is selected.
Train_samples_per_iteration: (DL) Specify the number of global training samples per MapReduce iteration. To specify one epoch, enter 0. To specify all available data (e.g., replicated training data), enter -1. To use the automatic values, enter -2. The default is -2.
Adaptive_rate: (DL) Check this checkbox to enable the adaptive learning rate (ADADELTA). This option is selected by default. If this option is enabled, the following parameters are ignored:
rate,rate_decay,rate_annealing,momentum_start,momentum_ramp,momentum_stable, andnesterov_accelerated_gradient.Input_dropout_ratio: (DL) Specify the input layer dropout ratio to improve generalization. Suggested values are 0.1 or 0.2. The range is >= 0 to <1 and the default value is 0.
L1: (DL) Specify the L1 regularization to add stability and improve generalization; sets the value of many weights to 0. The default value is 0.
L2: (DL) Specify the L2 regularization to add stability and improve generalization; sets the value of many weights to smaller values. The default value is 0.
Score_interval: (DL) Specify the shortest time interval (in seconds) to wait between model scoring. The default value is 5.
Score_training_samples: (DL) Specify the number of training set samples for scoring. To use all training samples, enter 0. The default value is 10000.
Score_validation_samples: (DL) (Requires selection from the Validation_Frame drop-down list) Specify the number of validation set samples for scoring. To use all validation set samples, enter 0. The default value is 0. This option is applicable to classification only.
Score_duty_cycle: (DL) Specify the maximum duty cycle fraction for scoring. A lower value results in more training and a higher value results in more scoring. The default value is 0.1.
Autoencoder: (DL) Check this checkbox to enable the Deep Learning autoencoder. This option is not selected by default. Note: This option requires a loss function other than CrossEntropy. If this option is enabled, use_all_factor_levels must be enabled.
Balance_classes: (GLM, GBM, DRF, DL, Naïve Bayes) Oversample the minority classes to balance the class distribution. This option is not selected by default. This option is only applicable for classification. Majority classes can be undersampled to satisfy the Max_after_balance_size parameter.
Max_confusion_matrix_size: (DRF, Naïve Bayes, GBM) Specify the maximum size (in number of classes) for confusion matrices to be printed in the Logs.
Max_hit_ratio_k: (DRF, Naïve Bayes) Specify the maximum number (top K) of predictions to use for hit ratio computation. Applicable to multi-class only. To disable, enter 0.
Link: (GLM) Select a link function (Identity, Family_Default, Logit, Log, or Inverse).
Alpha: (GLM) Specify the regularization distribution between L2 and L2. The default value is 0.5.
Lambda: (GLM) Specify the regularization strength. There is no default value.
Lambda_search: (GLM) Check this checkbox to enable lambda search, starting with lambda max. The given lambda is then interpreted as lambda min.
Rate: (DL) Specify the learning rate. Higher rates result in less stable models and lower rates result in slower convergence. The default value is 0.005. Not applicable if adaptive_rate is enabled.
Rate_annealing: (DL) Specify the learning rate annealing. The formula is rate/(1+rate_annealing value * samples). The default value is 10.000001. Not applicable if adaptive_rate is enabled.
Momentum_start: (DL) Specify the initial momentum at the beginning of training. A suggested value is 0.5. The default value is 0. Not applicable if adaptive_rate is enabled.
Momentum_ramp: (DL) Specify the number of training samples for increasing the momentum. The default value is 1000000. Not applicable if adaptive_rate is enabled.
Momentum_stable: DL Specify the final momentum value reached after the momentum_ramp training samples. Not applicable if adaptive_rate is enabled.
Nesterov_accelerated_gradient: (DL) Check this checkbox to use the Nesterov accelerated gradient. This option is recommended and selected by default. Not applicable is adaptive_rate is enabled.
Hidden_dropout_ratios: (DL) Specify the hidden layer dropout ratios to improve generalization. Specify one value per hidden layer, each value between 0 and 1 (exclusive). There is no default value. This option is applicable only if TanhwithDropout, RectifierwithDropout, or MaxoutWithDropout is selected from the Activation drop-down list.
Expert Options
Keep_cross_validation_splits: (DL) Check this checkbox to keep the cross-validation frames. This option is not selected by default.
Overwrite_with_best_model: (DL) Check this checkbox to overwrite the final model with the best model found during training. This option is selected by default.
Target_ratio_comm_to_comp: (DL) Specify the target ratio of communication overhead to computation. This option is only enabled for multi-node operation and if train_samples_per_iteration equals -2 (auto-tuning). The default value is 0.02.
Rho: (DL) Specify the adaptive learning rate time decay factor. The default value is 0.99. This option is only applicable if adaptive_rate is enabled.
Epsilon: (DL) Specify the adaptive learning rate time smoothing factor to avoid dividing by zero. The default value is 1.0E-8. This option is only applicable if adaptive_rate is enabled.
Max_W2: (DL) Specify the constraint for the squared sum of the incoming weights per unit (e.g., for Rectifier). The default value is infinity.
Initial_weight_distribution: (DL) Select the initial weight distribution (Uniform Adaptive, Uniform, or Normal). The default is Uniform Adaptive. If Uniform Adaptive is used, the initial_weight_scale parameter is not applicable.
Initial_weight_scale: (DL) Specify the initial weight scale of the distribution function for Uniform or Normal distributions. For Uniform, the values are drawn uniformly from initial weight scale. For Normal, the values are drawn from a Normal distribution with the standard deviation of the initial weight scale. The default value is 1.0. If Uniform Adaptive is selected as the initial_weight_distribution, the initial_weight_scale parameter is not applicable.
Classification_stop: (DL) (Applicable to discrete/categorical datasets only) Specify the stopping criterion for classification error fractions on training data. To disable this option, enter -1. The default value is 0.0.
Max_hit_ratio_k: (DL,)GLM (Classification only) Specify the maximum number (top K) of predictions to use for hit ratio computation (for multi-class only). To disable this option, enter 0. The default value is 10.
Regression_stop: (DL) (Applicable to real value/continuous datasets only) Specify the stopping criterion for regression error (MSE) on the training data. To disable this option, enter -1. The default value is 0.000001.
Diagnostics: (DL) Check this checkbox to compute the variable importances for input features (using the Gedeon method). For large networks, selecting this option can reduce speed. This option is selected by default.
Fast_mode: (DL) Check this checkbox to enable fast mode, a minor approximation in back-propagation. This option is selected by default.
Ignore_const_cols: Check this checkbox to ignore constant training columns, since no information can be gained from them. This option is selected by default.
Force_load_balance: (DL) Check this checkbox to force extra load balancing to increase training speed for small datasets and use all cores. This option is selected by default.
Single_node_mode: (DL) Check this checkbox to force H2O to run on a single node for fine-tuning of model parameters. This option is not selected by default.
Replicate_training_data: (DL) Check this checkbox to replicate the entire training dataset on every node for faster training on small datasets. This option is not selected by default. This option is only applicable for clouds with more than one node.
Shuffle_training_data: (DL) Check this checkbox to shuffle the training data. This option is recommended if the training data is replicated and the value of train_samples_per_iteration is close to the number of nodes times the number of rows. This option is not selected by default.
Missing_values_handling: (DL) Select how to handle missing values (Skip or MeanImputation). The default value is MeanImputation.
Quiet_mode: (DL) Check this checkbox to display less output in the standard output. This option is not selected by default.
Sparse: (DL) Check this checkbox to use sparse iterators for the input layer. This option is not selected by default as it rarely improves performance.
Col_major: (DL) Check this checkbox to use a column major weight matrix for the input layer. This option can speed up forward propagation but may reduce the speed of backpropagation. This option is not selected by default.
Average_activation: (DL) Specify the average activation for the sparse autoencoder. The default value is 0. If Rectifier is selected as the Activation type, this value must be positive. For Tanh, the value must be in (-1,1).
Sparsity_beta: (DL) Specify the sparsity regularization. The default value is 0.
Max_categorical_features: (DL) Specify the maximum number of categorical features enforced via hashing. The default is unlimited.
Reproducible: (DL) To force reproducibility on small data, check this checkbox. If this option is enabled, the model takes more time to generate, since it uses only one thread.
Export_weights_and_biases: (DL) To export the neural network weights and biases as H2O frames, check this checkbox.
Class_sampling_factors: (GLM, DRF, Naïve Bayes), GBM, DL) Specify the per-class (in lexicographical order) over/under-sampling ratios. By default, these ratios are automatically computed during training to obtain the class balance. There is no default value. This option is only applicable for classification problems and when Balance_Classes is enabled.
Seed: (K-Means, GBM, DL, DRF) Specify the random number generator (RNG) seed for algorithm components dependent on randomization. The seed is consistent for each H2O instance so that you can create models with the same starting conditions in alternative configurations.
Prior: (GLM) Specify prior probability for y ==1. Use this parameter for logistic regression if the data has been sampled and the mean of response does not reflect reality. The default value is -1.
Max_active_predictors: (GLM) Specify the maximum number of active predictors during computation. This value is used as a stopping criterium to prevent expensive model building with many predictors.
Viewing Models
Click the Assist Me! button, then click the getModels link, or enter getModels in the cell in CS mode and press Ctrl+Enter. A list of available models displays.
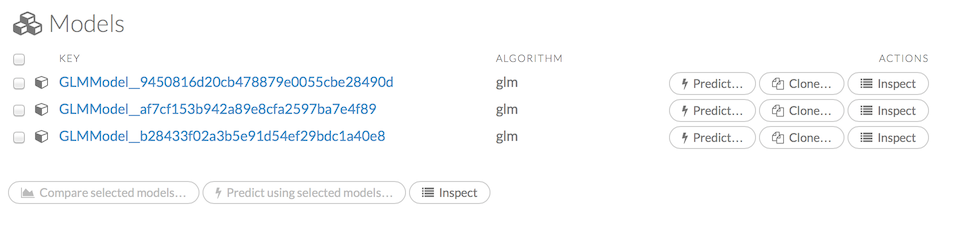
To view all current models, you can also click the Model menu and click List All Models.
To inspect a model, check its checkbox then click the Inspect button, or click the Inspect button to the right of the model name.
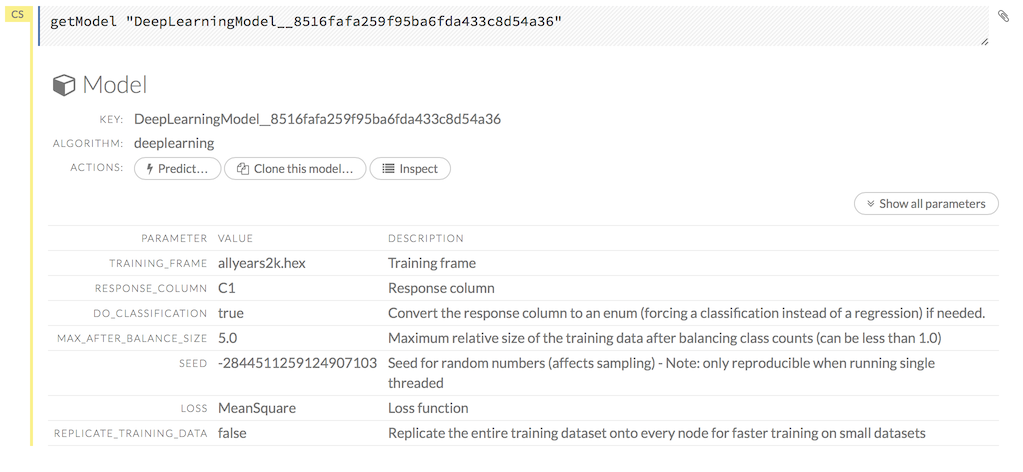
A summary of the model’s parameters displays. To display more details, click the Show All Parameters button.
NOTE: The Clone this model… button will be supported in a future version.
To delete a model, click the Delete button.
To generate a POJO to be able to use the model outside of H2O, click the Preview POJO button.
To learn how to make predictions, continue to the next section.
… Making Predictions
After creating your model, click the key link for the model, then click the Predict button. Select the model to use in the prediction from the drop-down Model: menu and the data frame to use in the prediction from the drop-down Frame menu, then click the Predict button.
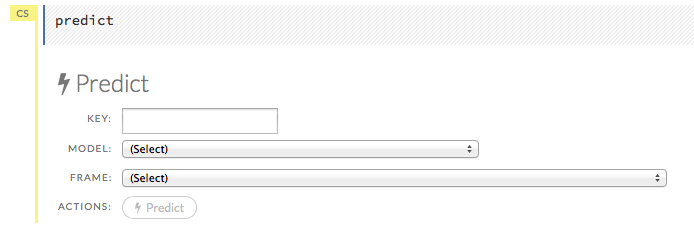
Viewing Predictions
Click the Assist Me! button, then click the getPredictions link, or enter getPredictions in the cell in CS mode and press Ctrl+Enter. A list of the stored predictions displays.
To view a prediction, click the View button to the right of the model name.
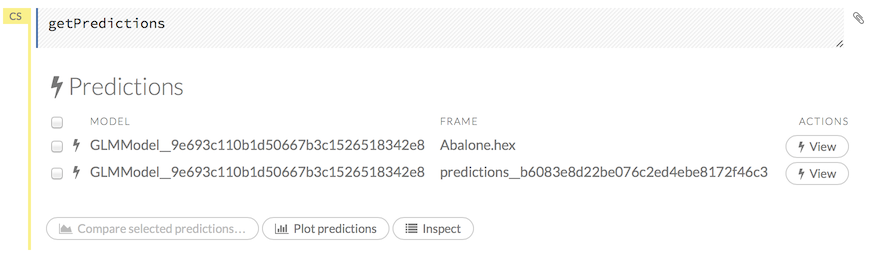
You can also view predictions by clicking the drop-down Score menu and selecting List All Predictions.
Viewing Frames
To view a specific frame, click the “Key” link for the specified frame, or enter getFrameSummary "FrameName" in a cell in CS mode (where FrameName is the name of a frame, such as allyears2k.hex.
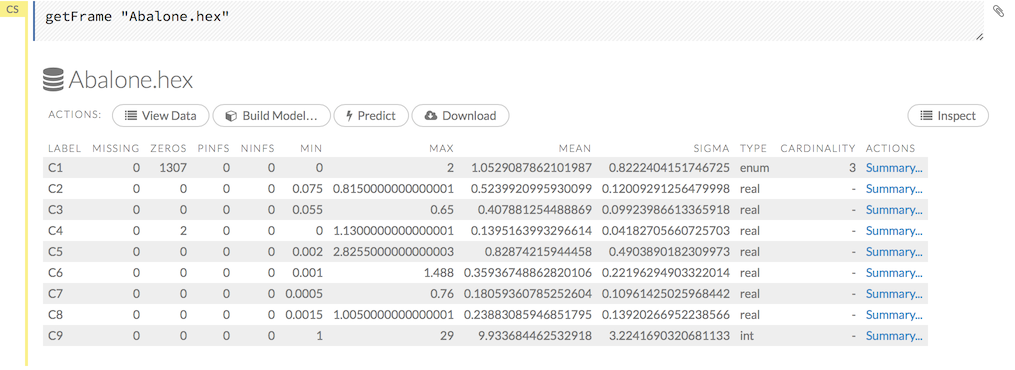
From the getFrameSummary cell, you can:
- view a truncated list of the rows in the data frame by clicking the View Data button
- split the dataset by clicking the Split… button
- view the columns, data, and factors in more detail or plot a graph by clicking the Inspect button
- create a model by clicking the Build Model button
- make a prediction based on the data by clicking the Predict button
- download the data as a .csv file by clicking the Download button
- view the characteristics or domain of a specific column by clicking the Summary link
When you view a frame, you can “drill-down” to the necessary level of detail (such as a specific column or row) using the Inspect button or by clicking the links. The following screenshot displays the results of clicking the Inspect button for a frame.
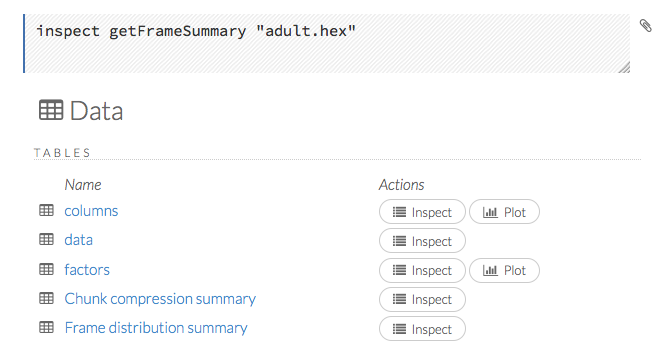
This screenshot displays the results of clicking the columns link.
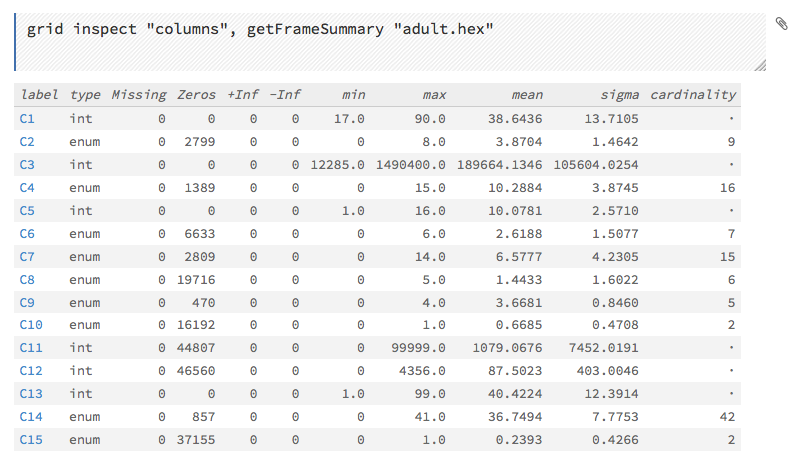
To view all frames, click the Assist Me! button, then click the getFrames link, or enter getFrames in the cell in CS mode and press Ctrl+Enter. You can also view all current frames by clicking the drop-down Data menu and selecting List All Frames.
A list of the current frames in H2O displays that includes the following information for each frame:
- Link to the frame (the “key”)
- Number of rows and columns
- Size
For parsed data, the following information displays:
- Link to the .hex file
The Build Model, Predict, and Inspect buttons
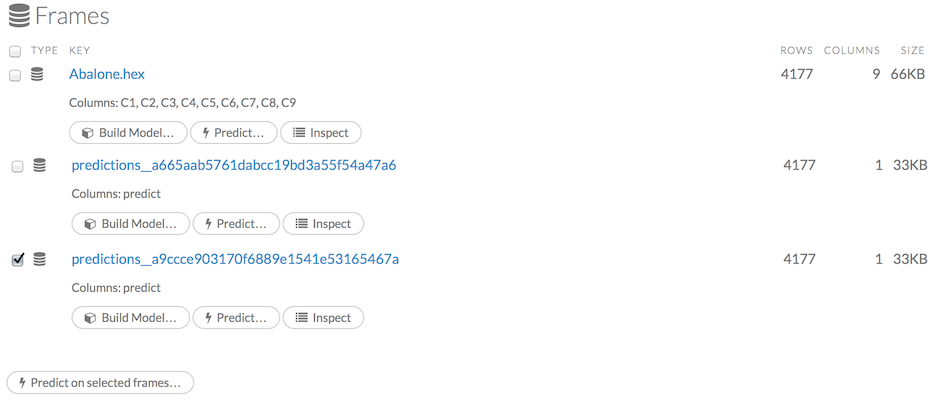
To make a prediction, check the checkboxes for the frames you want to use to make the prediction, then click the Predict on Selected Frames button.
Splitting Frames
Datasets can be split within Flow for use in model training and testing.
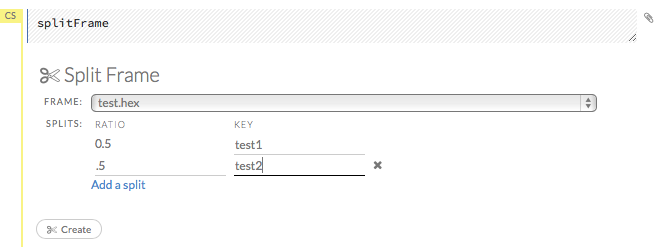
- To split a frame, click the Assist Me button, then click splitFrame. Note: You can also click the drop-down Data menu and select Split Frame….
- From the drop-down Frame: list, select the frame to split.
In the second Ratio entry field, specify the fractional value to determine the split. The first Ratio field is automatically calculated based on the values entered in the second Ratio field.
Note: Only fractional values between 0 and 1 are supported (for example, enter
.5to split the frame in half). The total sum of the ratio values must equal one. H2O automatically adjusts the ratio values to equal one; if unsupported values are entered, an error displays.- In the Key entry field, specify a name for the new frame.
- (Optional) To add another split, click the Add a split link. To remove a split, click the
Xto the right of the Key entry field. - Click the Create button.
Creating Frames
To create a frame with a large amount of random data (for example, to use for testing), click the drop-down Admin menu, then select Create Synthetic Frame. Customize the frame as needed, then click the Create button to create the frame.
Plotting Frames
To create a plot from a frame, click the Inspect button, then click the Plot button.
Select the type of plot (point, path, or rect) from the drop-down Type menu, then select the x-axis and y-axis from the following options:
- label
- type
- missing
- zeros
- +Inf
- -Inf
- min
- max
- mean
- sigma
- cardinality
Select one of the above options from the drop-down Color menu to display the specified data in color, then click the Plot button to plot the data.
Note: Because H2O stores enums internally as numeric then maps the integers to an array of strings, any min, max, or mean values for categorical columns are not meaningful and should be ignored. Displays for categorical data will be modified in a future version of H2O.
… Using Flows
You can use and modify flows in a variety of ways:
- Clips allow you to save single cells
- Outlines display summaries of your workflow
- Flows can be saved, duplicated, loaded, or downloaded
Using Clips
Clips enable you to save cells containing your workflow for later reuse. To save a cell as a clip, click the paperclip icon to the right of the cell (highlighted in the red box in the following screenshot).

To use a clip in a workflow, click the “Clips” tab in the sidebar on the right.
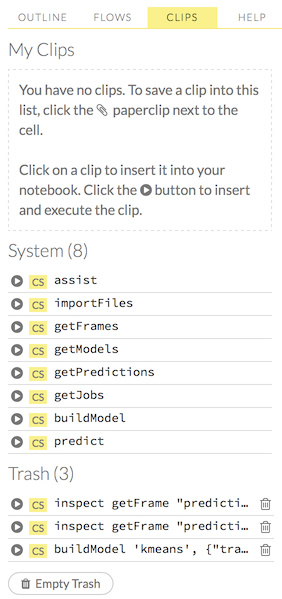
All saved clips, including the default system clips (such as assist, importFiles, and predict), are listed. Clips you have created are listed under the “My Clips” heading. To select a clip to insert, click the circular button to the left of the clip name. To delete a clip, click the trashcan icon to right of the clip name.
NOTE: The default clips listed under “System” cannot be deleted.
Deleted clips are stored in the trash. To permanently delete all clips in the trash, click the Empty Trash button.
NOTE: Saved data, including flows and clips, are persistent as long as the same IP address is used for the cluster. If a new IP is used, previously saved flows and clips are not available.
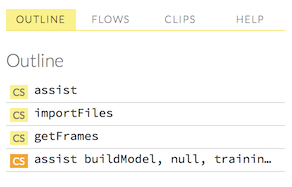
Viewing Outlines
The “Outline” tab in the sidebar displays a brief summary of the cells currently used in your flow; essentially, a command history.
- To jump to a specific cell, click the cell description.
To delete a cell, select it and press the X key on your keyboard.
Saving Flows
- Finding Saved Flows on your Disk
- Saving Flows on a Hadoop cluster
- Copying Flows
- Downloading Flows
- Loading Flows
You can save your flow for later reuse. To save your flow as a notebook, click the “Save” button (the first button in the row of buttons below the flow name), or click the drop-down “Flow” menu and select “Save.” To enter a custom name for the flow, click the default flow name (“Untitled Flow”) and type the desired flow name. A pencil icon indicates where to enter the desired name.

To confirm the name, click the checkmark to the right of the name field.
To reuse a saved flow, click the “Flows” tab in the sidebar, then click the flow name. To delete a saved flow, click the trashcan icon to the right of the flow name.
Finding Saved Flows on your Disk
By default, flows are saved to the h2oflows directory underneath your home directory. The directory where flows are saved is printed to stdout:
03-20 14:54:20.945 172.16.2.39:54323 95667 main INFO: Flow dir: '/Users/<UserName>/h2oflows'
To back up saved flows, copy this directory to your preferred backup location.
To specify a different location for saved flows, use the command-line argument -flow_dir when launching H2O:
java -jar h2o.jar -flow_dir /<New>/<Location>/<For>/<Saved>/<Flows>
where /<New>/<Location>/<For>/<Saved>/<Flows> represents the specified location. If the directory does not exist, it will be created the first time you save a flow.
Saving Flows on a Hadoop cluster
Note: If you are running H2O Flow on a Hadoop cluster, H2O will try to find the HDFS home directory to use as the default directory for flows. If the HDFS home directory is not found, flows cannot be saved unless a directory is specified while launching using -flow_dir:
hadoop jar h2odriver.jar -nodes 1 -mapperXmx 1g -output hdfsOutputDirName -flow_dir hdfs:///<Saved>/<Flows>/<Location>
The location specified in flow_dir may be either an hdfs or regular filesystem directory. If the directory does not exist, it will be created the first time you save a flow.
Copying Flows
To create a copy of the current flow, select the Flow menu, then click Make a Copy. The name of the current flow changes to “Copy of
Downloading Flows
After saving a flow as a notebook, click the Flow menu, then select Download…. A new window opens and the saved flow is downloaded to the default downloads folder on your computer. The file is exported as <filename>.flow, where <filename> is the name specified when the flow was saved.
Caution: You must have an active internet connection to download flows.
Loading Flows
To load a saved flow, click the Flows tab in the sidebar at the right. In the pop-up confirmation window that appears, select Load Notebook, or click Cancel to return to the current flow.
After clicking Load Notebook, the saved flow is loaded.
To load an exported flow, click the Flow menu and select Open…. In the pop-up window that appears, click the Choose File button and select the exported flow, then click the Open button.
Notes:
- Only exported flows using the default .flow filetype are supported. Other filetypes will not open.
- If the current notebook has the same name as the selected file, a pop-up confirmation appears to confirm that the current notebook should be overwritten.
…Troubleshooting Flow
- Viewing Cluster Status
- Viewing CPU Status (Water Meter)
- Viewing Logs
- Downloading Logs
- Viewing Stack Trace Information
- Viewing Network Test Results
- Accessing the Profiler
- Viewing the Timeline
- Shutting Down H2O
To troubleshoot issues in Flow, use the Admin menu. The Admin menu allows you to check the status of the cluster, view a timeline of events, and view or download logs for issue analysis.
NOTE: To view the current version, click the Help menu, then click About.
Viewing Cluster Status
Click the Admin menu, then select Cluster Status. A summary of the status of the cluster (also known as a cloud) displays, which includes the same information:
- Cluster health
- Whether all nodes can communicate (consensus)
Whether new nodes can join (locked/unlocked)
Note: After you submit a job to H2O, the cluster does not accept new nodes.
- H2O version
- Number of used and available nodes
When the cluster was created
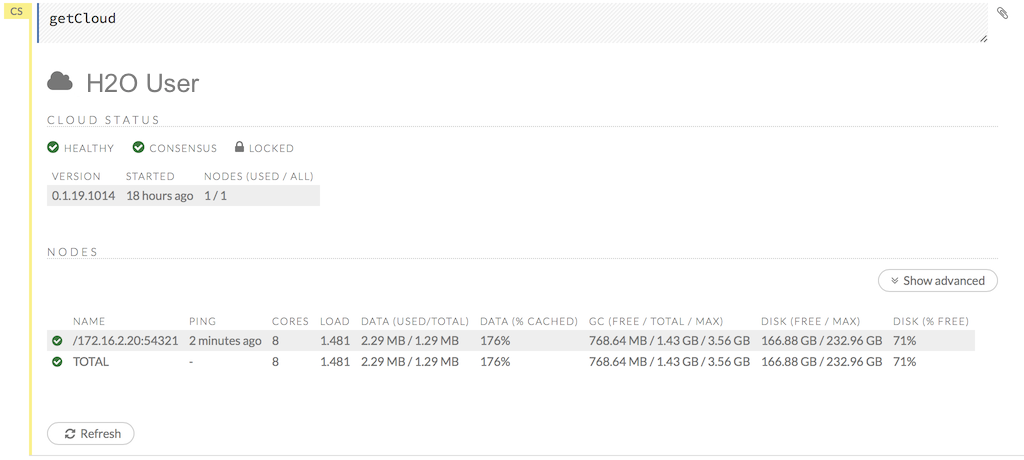
The following information displays for each node:
- IP address (name)
- Time of last ping
- Number of cores
- Load
- Amount of data (used/total)
- Percentage of cached data
- GC (free/total/max)
- Amount of disk space in GB (free/max)
- Percentage of free disk space
To view more information, click the Show Advanced button.
Viewing CPU Status (Water Meter)
To view the current CPU usage, click the Admin menu, then click Water Meter (CPU Meter). A new window opens, displaying the current CPU use statistics.
Viewing Logs
To view the logs for troubleshooting, click the Admin menu, then click Inspect Log.
To view the logs for a specific node, select it from the drop-down Select Node menu.
Downloading Logs
To download the logs for further analysis, click the Admin menu, then click Download Log. A new window opens and the logs download to your default download folder. You can close the new window after downloading the logs. Send the logs to support@h2o.ai for issue resolution.
Viewing Stack Trace Information
To view the stack trace information, click the Admin menu, then click Stack Trace.
To view the stack trace information for a specific node, select it from the drop-down Select Node menu.
Viewing Network Test Results
To view network test results, click the Admin menu, then click Network Test.
Accessing the Profiler
The Profiler looks across the cluster to see where the same stack trace occurs, and can be helpful for identifying what the currently used CPU is doing. To view the profiler, click the Admin menu, then click Profiler.
To view the profiler information for a specific node, select it from the drop-down Select Node menu.
Viewing the Timeline
To view a timeline of events in Flow, click the Admin menu, then click Timeline. The following information displays for each event:
- Time of occurrence (HH:MM:SS:MS)
- Number of nanoseconds for duration
- Originator of event (“who”)
- I/O type
- Event type
Number of bytes sent & received
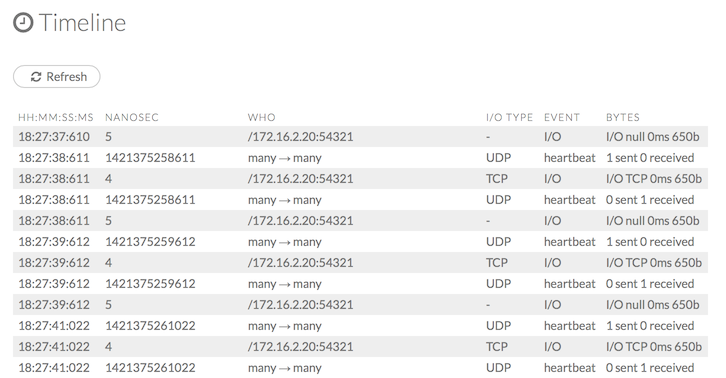
To obtain the most recent information, click the Refresh button.
Shutting Down H2O
To shut down H2O, click the Admin menu, then click Shut Down. A Shut down complete message displays in the upper right when the cluster has been shut down.
Data Science Algorithms
This document describes how to define the models, interpret the model, the algorithm itself, and an FAQ.
Commonalities
Missing Value Handling for Training
If missing values are found in the validation frame during model training or during the scoring process for creating predictions, the missing values are automatically imputed.
If the missing values are found during POJO scoring, the answer is converted to NaN.
K-Means
Introduction
K-Means falls in the general category of clustering algorithms.
Defining a K-Means Model
Destination_key: (Optional) Enter a custom name for the model to use as a reference. By default, H2O automatically generates a destination key.
Training_frame: (Required) Select the dataset used to build the model. NOTE: If you click the Build a model button from the
Parsecell, the training frame is entered automatically.Validation_frame: (Optional) Select the dataset used to evaluate the accuracy of the model.
Ignored_columns: (Optional) Click the plus sign next to a column name to add it to the list of columns excluded from the model. To add all columns, click the Add all button. To remove a column from the list of ignored columns, click the X next to the column name. To remove all columns from the list of ignored columns, click the Clear all button.
Ignore_const_cols: (Optional) Check this checkbox to ignore constant training columns, since no information can be gained from them. This option is selected by default.
Score_each_iteration: (Optional) Check this checkbox to score during each iteration of the model training.
K: Specify the number of clusters. The default is 1.
User_points: Specify a vector of initial cluster centers.
Max_iterations: Specify the maximum number of training iterations. The default is 1000.
Init: Select the initialization mode. The options are Random, Furthest, PlusPlus, or User. Note: If PlusPlus is selected, the initial Y matrix is chosen by the final cluster centers from the K-Means PlusPlus algorithm.
Standardize: To standardize the numeric columns to have mean of zero and unit variance, check this checkbox. Standardization is highly recommended; if you do not use standardization, the results can include components that are dominated by variables that appear to have larger variances relative to other attributes as a matter of scale, rather than true contribution. This option is selected by default.
Seed: Specify the random number generator (RNG) seed for algorithm components dependent on randomization. The seed is consistent for each H2O instance so that you can create models with the same starting conditions in alternative configurations.
Interpreting a K-Means Model
By default, the following output displays:
- A graph of the scoring history (number of iterations vs. average within the cluster’s sum of squares)
- Output (model category, validation metrics if applicable, and centers std)
- Model Summary (number of clusters, number of categorical columns, number of iterations, avg. within sum of squares, avg. sum of squares, avg. between the sum of squares)
- Scoring history (number of iterations, avg. change of standardized centroids, avg. within cluster sum of squares)
- Training metrics (model name, checksum name, frame name, frame checksum name, description if applicable, model category, duration in ms, scoring time, predictions, MSE, avg. within sum of squares, avg. between sum of squares)
- Centroid statistics (centroid number, size, within sum of squares)
- Cluster means (centroid number, column)
K-Means randomly chooses starting points and converges to a local minimum of centroids. The number of clusters is arbitrary, and should be thought of as a tuning parameter. The output is a matrix of the cluster assignments and the coordinates of the cluster centers in terms of the originally chosen attributes. Your cluster centers may differ slightly from run to run as this problem is Non-deterministic Polynomial-time (NP)-hard.
FAQ
How does the algorithm handle missing values during training?
Missing values are automatically imputed by the column mean.
How does the algorithm handle missing values during testing?
Missing values are automatically imputed by the column mean of the training data.
Does it matter if the data is sorted?
No.
Should data be shuffled before training?
No.
What if there are a large number of columns?
K-Means suffers from the curse of dimensionality: all points are roughly at the same distance from each other in high dimensions, making the algorithm less and less useful.
What if there are a large number of categorical factor levels?
This can be problematic, as categoricals are one-hot encoded on the fly, which can lead to the same problem as datasets with a large number of columns.
K-Means Algorithm
The number of clusters \(K\) is user-defined and is determined a priori.
Choose \(K\) initial cluster centers \(m_{k}\) according to one of the following:
Randomization: Choose \(K\) clusters from the set of \(N\) observations at random so that each observation has an equal chance of being chosen.
Plus Plus
a. Choose one center \(m_{1}\) at random.
Calculate the difference between \(m_{1}\) and each of the remaining \(N-1\) observations \(x_{i}\). \(d(x_{i}, m_{1}) = ||(x_{i}-m_{1})||^2\)
Let \(P(i)\) be the probability of choosing \(x_{i}\) as \(m_{2}\). Weight \(P(i)\) by \(d(x_{i}, m_{1})\) so that those \(x_{i}\) furthest from \(m_{2}\) have a higher probability of being selected than those \(x_{i}\) close to \(m_{1}\).
Choose the next center \(m_{2}\) by drawing at random according to the weighted probability distribution.
Repeat until \(K\) centers have been chosen.
Furthest
a. Choose one center \(m_{1}\) at random.
Calculate the difference between \(m_{1}\) and each of the remaining \(N-1\) observations \(x_{i}\). \(d(x_{i}, m_{1}) = ||(x_{i}-m_{1})||^2\)
Choose \(m_{2}\) to be the \(x_{i}\) that maximizes \(d(x_{i}, m_{1})\).
Repeat until \(K\) centers have been chosen.
Once \(K\) initial centers have been chosen calculate the difference between each observation \(x_{i}\) and each of the centers \(m_{1},...,m_{K}\), where difference is the squared Euclidean distance taken over \(p\) parameters.
\(d(x_{i}, m_{k})=\) \(\sum_{j=1}^{p}(x_{ij}-m_{k})^2=\) \(\lVert(x_{i}-m_{k})\rVert^2\)
Assign \(x_{i}\) to the cluster \(k\) defined by \(m_{k}\) that minimizes \(d(x_{i}, m_{k})\)
When all observations \(x_{i}\) are assigned to a cluster calculate the mean of the points in the cluster.
\(\bar{x}(k)=\lbrace\bar{x_{i1}},…\bar{x_{ip}}\rbrace\)
Set the \(\bar{x}(k)\) as the new cluster centers \(m_{k}\). Repeat steps 2 through 5 until the specified number of max iterations is reached or cluster assignments of the \(x_{i}\) are stable.
References
Xiong, Hui, Junjie Wu, and Jian Chen. “K-means Clustering Versus Validation Measures: A Data- distribution Perspective.” Systems, Man, and Cybernetics, Part B: Cybernetics, IEEE Transactions on 39.2 (2009): 318-331.
GLM
Introduction
Generalized Linear Models (GLM) estimate regression models for outcomes following exponential distributions. In addition to the Gaussian (i.e. normal) distribution, these include Poisson, binomial, and gamma distributions. Each serves a different purpose, and depending on distribution and link function choice, can be used either for prediction or classification.
The GLM suite includes:
- Gaussian regression
- Poisson regression
- Binomial regression
- Gamma regression
Defining a GLM Model
Destination_key: (Optional) Enter a custom name for the model to use as a reference. By default, H2O automatically generates a destination key.
Training_frame: (Required) Select the dataset used to build the model. NOTE: If you click the Build a model button from the
Parsecell, the training frame is entered automatically.Validation_frame: (Optional) Select the dataset used to evaluate the accuracy of the model.
Ignored_columns: (Optional) Click the plus sign next to a column name to add it to the list of columns excluded from the model. To add all columns, click the Add all button. To remove a column from the list of ignored columns, click the X next to the column name. To remove all columns from the list of ignored columns, click the Clear all button.
Ignore_const_cols: Check this checkbox to ignore constant training columns, since no information can be gained from them. This option is selected by default.
Response_column: (Required) Select the column to use as the independent variable.
Family: Select the model type (Gaussian, Binomial, Poisson, or Gamma).
Solver: Select the solver to use (IRLSM, L_BFGS, or auto). IRLSM is fast on on problems with small number of predictors and for lambda-search with L1 penalty, while L_BFGS scales better for datasets with many columns. The default is IRLSM.
Alpha: Specify the regularization distribution between L2 and L2. The default value is 0.5.
Lambda: Specify the regularization strength. The default value is data dependent.
Lambda_search: Check this checkbox to enable lambda search, starting with lambda max. The given lambda is then interpreted as lambda min.
Standardize: To standardize the numeric columns to have a mean of zero and unit variance, check this checkbox. Standardization is highly recommended; if you do not use standardization, the results can include components that are dominated by variables that appear to have larger variances relative to other attributes as a matter of scale, rather than true contribution. This option is selected by default.
Beta constraints: To use beta constraints, select a dataset from the drop-down menu. The selected frame is used to constraint the coefficient vector to provide upper and lower bounds.
Max_confusion_matrix_size: Specify the maximum size (number of classes) for the confusion matrices printed in the logs.
Max_hits_ratio_k: Specify the maximum number (top K) of predictions to use for hit ratio computation. Applicable to multi-class only. To disable, enter
0.Max_iterations: Specify the number of training iterations. The default is 50.
Beta_epsilon: Specify the beta epsilon value. If the L1 normalization of the current beta change is below this threshold, consider using convergence.
Link: Select a link function (Identity, Family_Default, Logit, Log, or Inverse).
Prior: Specify prior probability for y ==1. Use this parameter for logistic regression if the data has been sampled and the mean of response does not reflect reality. The default value is 0.
Max_active_\predictors: Specify the maximum number of active predictors during computation. This value is used as a stopping criterium to prevent expensive model building with many predictors.
Interpreting a GLM Model
By default, the following output displays:
- A graph of the normalized coefficient magnitudes
- Output (model category, model summary, scoring history, training metrics, validation metrics, best lambda, threshold, residual deviance, null deviance, residual degrees of freedom, null degrees of freedom, AIC, AUC, binomial, rank)
- Coefficients
- Coefficient magnitudes
FAQ
How does the algorithm handle missing values during training?
GLM skips rows with missing values.
How does the algorithm handle missing values during testing?
GLM will predict Double.NaN for rows containg missing values.
What happens if the response has missing values?
It is handled properly, but verify the results are correct.
What happens during prediction if the new sample has categorical levels not seen in training?
It will predict Double.NaN.
Does it matter if the data is sorted?
No.
Should data be shuffled before training?
No.
How does the algorithm handle highly imbalanced data in a response column?
GLM does not require special handling for imbalanced data.
What if there are a large number of columns?
IRLS will get quadratically slower with the number of columns. Try L-BFGS for datasets with more than 5-10 thousand columns.
What if there are a large number of categorical factor levels?
GLM internally one-hot encodes the categorical factor levels; the same limitations as with a high column count will apply.
GLM Algorithm
Following the definitive text by P. McCullagh and J.A. Nelder (1989) on the generalization of linear models to non-linear distributions of the response variable Y, H2O fits GLM models based on the maximum likelihood estimation via iteratively reweighed least squares.
Let \(y_{1},…,y_{n}\) be n observations of the independent, random response variable \(Y_{i}\).
Assume that the observations are distributed according to a function from the exponential family and have a probability density function of the form:
\(f(y_{i})=exp[\frac{y_{i}\theta_{i} - b(\theta_{i})}{a_{i}(\phi)} + c(y_{i}; \phi)]\) where \(\theta\) and \(\phi\) are location and scale parameters, and \(\: a_{i}(\phi), \:b_{i}(\theta_{i}),\: c_{i}(y_{i}; \phi)\) are known functions.
\(a_{i}\) is of the form \(\:a_{i}=\frac{\phi}{p_{i}}; p_{i}\) is a known prior weight.
When \(Y\) has a pdf from the exponential family:
\(E(Y_{i})=\mu_{i}=b^{\prime}\) \(var(Y_{i})=\sigma_{i}^2=b^{\prime\prime}(\theta_{i})a_{i}(\phi)\)
Let \(g(\mu_{i})=\eta_{i}\) be a monotonic, differentiable transformation of the expected value of \(y_{i}\). The function \(\eta_{i}\) is the link function and follows a linear model.
\(g(\mu_{i})=\eta_{i}=\mathbf{x_{i}^{\prime}}\beta\)
When inverted: \(\mu=g^{-1}(\mathbf{x_{i}^{\prime}}\beta)\)
Maximum Likelihood Estimation
For an initial rough estimate of the parameters \(\hat{\beta}\), use the estimate to generate fitted values: \(\mu_{i}=g^{-1}(\hat{\eta_{i}})\)
Let \(z\) be a working dependent variable such that \(z_{i}=\hat{\eta_{i}}+(y_{i}-\hat{\mu_{i}})\frac{d\eta_{i}}{d\mu_{i}}\),
where \(\frac{d\eta_{i}}{d\mu_{i}}\) is the derivative of the link function evaluated at the trial estimate.
Calculate the iterative weights: \(w_{i}=\frac{p_{i}}{[b^{\prime\prime}(\theta_{i})\frac{d\eta_{i}}{d\mu_{i}}^{2}]}\)
Where \(b^{\prime\prime}\) is the second derivative of \(b(\theta_{i})\) evaluated at the trial estimate.
Assume \(a_{i}(\phi)\) is of the form \(\frac{\phi}{p_{i}}\). The weight \(w_{i}\) is inversely proportional to the variance of the working dependent variable \(z_{i}\) for current parameter estimates and proportionality factor \(\phi\).
Regress \(z_{i}\) on the predictors \(x_{i}\) using the weights \(w_{i}\) to obtain new estimates of \(\beta\). \(\hat{\beta}=(\mathbf{X}^{\prime}\mathbf{W}\mathbf{X})^{-1}\mathbf{X}^{\prime}\mathbf{W}\mathbf{z}\)
Where \(\mathbf{X}\) is the model matrix, \(\mathbf{W}\) is a diagonal matrix of \(w_{i}\), and \(\mathbf{z}\) is a vector of the working response variable \(z_{i}\).
This process is repeated until the estimates \(\hat{\beta}\) change by less than the specified amount.
Cost of computation
H2O can process large data sets because it relies on parallel processes. Large data sets are divided into smaller data sets and processed simultaneously and the results are communicated between computers as needed throughout the process.
In GLM, data are split by rows but not by columns, because the predicted Y values depend on information in each of the predictor variable vectors. If O is a complexity function, N is the number of observations (or rows), and P is the number of predictors (or columns) then
\(Runtime\propto p^3+\frac{(N*p^2)}{CPUs}\)
Distribution reduces the time it takes an algorithm to process because it decreases N.
Relative to P, the larger that (N/CPUs) becomes, the more trivial p becomes to the overall computational cost. However, when p is greater than (N/CPUs), O is dominated by p.
\(Complexity = O(p^3 + N*p^2)\)
References
Breslow, N E. “Generalized Linear Models: Checking Assumptions and Strengthening Conclusions.” Statistica Applicata 8 (1996): 23-41.
Frome, E L. “The Analysis of Rates Using Poisson Regression Models.” Biometrics (1983): 665-674.
Snee, Ronald D. “Validation of Regression Models: Methods and Examples.” Technometrics 19.4 (1977): 415-428.
DRF
Introduction
Distributed Random Forest (DRF) is a powerful classification tool. When given a set of data, DRF generates a forest of classification trees, rather than a single classification tree. Each of these trees is a weak learner built on a subset of rows and columns. More trees will reduce the variance. The classification from each H2O tree can be thought of as a vote; the most votes determines the classification.
Defining a DRF Model
Destination_key: (Optional) Enter a custom name for the model to use as a reference. By default, H2O automatically generates a destination key.
Training_frame: (Required) Select the dataset used to build the model. NOTE: If you click the Build a model button from the
Parsecell, the training frame is entered automatically.Validation_frame: (Optional) Select the dataset used to evaluate the accuracy of the model.
Ignored_columns: (Optional) Click the plus sign next to a column name to add it to the list of columns excluded from the model. To add all columns, click the Add all button. To remove a column from the list of ignored columns, click the X next to the column name. To remove all columns from the list of ignored columns, click the Clear all button.
Ignore_const_cols: Check this checkbox to ignore constant training columns, since no information can be gained from them. This option is selected by default.
Score_each_iteration: (Optional) Check this checkbox to score during each iteration of the model training.
Response_column: (Required) Select the column to use as the independent variable.
Ntrees: Specify the number of trees. The default value is 50.
Max_depth: Specify the maximum tree depth. The default value is 5.
Min_rows: Specify the minimum number of observations for a leaf (
nodesizein R). The default value is 10.Nbins: Specify the number of bins for the histogram. The default value is 20.
Mtries: Specify the columns to randomly select at each level. To use the square root of the columns, enter
-1. The default value is -1.Sample_rate: Specify the sample rate. The range is 0 to 1.0 and the default value is 0.632.
Build_tree_one_node: To run on a single node, check this checkbox. This is suitable for small datasets as there is no network overhead but fewer CPUs are used. The default setting is disabled.
Balance_classes: Oversample the minority classes to balance the class distribution. This option is not selected by default. This option is only applicable for classification. Majority classes can be undersampled to satisfy the Max_after_balance_size parameter.
Max_confusion_matrix_size: Specify the maximum size (in number of classes) for confusion matrices to be printed in the Logs.
Max_hit_ratio_k: Specify the maximum number (top K) of predictions to use for hit ratio computation. Applicable to multi-class only. To disable, enter 0.
Class_sampling_factors: Specify the per-class (in lexicographical order) over/under-sampling ratios. By default, these ratios are automatically computed during training to obtain the class balance. There is no default value.
Seed: Specify the random number generator (RNG) seed for algorithm components dependent on randomization. The seed is consistent for each H2O instance so that you can create models with the same starting conditions in alternative configurations.
Interpreting a DRF Model
By default, the following output displays:
- Model parameters (hidden)
- A graph of the scoring history (number of trees vs. training MSE)
- A graph of the ROC curve (TPR vs. FPR)
- A graph of the variable importances
- Output (model category, validation metrics, initf)
- Model summary (number of trees, min. depth, max. depth, mean depth, min. leaves, max. leaves, mean leaves)
- Scoring history in tabular format
- Training metrics (model name, checksum name, frame name, frame checksum name, description, model category, duration in ms, scoring time, predictions, MSE, R2, logloss, AUC, GINI)
- Training metrics for thresholds (thresholds, F1, F2, F0Points, Accuracy, Precision, Recall, Specificity, Absolute MCC, min. per-class accuracy, TNS, FNS, FPS, TPS, IDX)
- Maximum metrics (metric, threshold, value, IDX)
- Variable importances in tabular format
FAQ
How does the algorithm handle missing values during training?
Missing values do not alter the tree building in any way (i.e., they are not counted as a point when computing means or errors). Rows containing missing values do affect tree building, but the missing values don’t change the split-point of the column they are in.
How does the algorithm handle missing values during testing?
During scoring, missing values “always go left” at any decision point in a tree. Due to dynamic binning in DRF, a row with a missing value typically ends up in the “leftmost bin” - with other outliers.
What happens if the response has missing values?
No errors will occur, but nothing will be learned from rows containing missing the response.
Does it matter if the data is sorted?
No.
Should data be shuffled before training?
No.
How does the algorithm handle highly imbalanced data in a response column?
Specify
balance_classes,class_sampling_factorsandmax_after_balance_sizeto control over/under-sampling.What if there are a large number of columns?
DRFs are best for datasets with fewer than a few thousand columns.
What if there are a large number of categorical factor levels?
Large numbers of categoricals are handled very efficiently - there is never any one-hot encoding.
DRF Algorithm
References
Naïve Bayes
- Introduction
- Defining a Naïve Bayes Model
- Interpreting a Naïve Bayes Model
- FAQ
- Naïve Bayes Algorithm
- References
Introduction
Naïve Bayes (NB) is a classification algorithm that relies on strong assumptions of the independence of covariates in applying Bayes Theorem. NB models are commonly used as an alternative to decision trees for classification problems.
Defining a Naïve Bayes Model
Destination_key: (Optional) Enter a custom name for the model to use as a reference. By default, H2O automatically generates a destination key.
Training_frame: (Required) Select the dataset used to build the model. NOTE: If you click the Build a model button from the
Parsecell, the training frame is entered automatically.Validation_frame: (Optional) Select the dataset used to evaluate the accuracy of the model.
Ignored_columns: (Optional) Click the plus sign next to a column name to add it to the list of columns excluded from the model. To add all columns, click the Add all button. To remove a column from the list of ignored columns, click the X next to the column name. To remove all columns from the list of ignored columns, click the Clear all button.
Ignore_const_cols: Check this checkbox to ignore constant training columns, since no information can be gained from them. This option is selected by default.
Response_column: (Required) Select the column to use as the independent variable.
Laplace: Specify the Laplace smoothing parameter. The default value is 0.
Min_sdev: Specify the minimum standard deviation to use for observations without enough data. The default value is 0.001.
Eps_sdev: Specify the threshold for standard deviation. If this threshold is not met, the min_sdev value is used. The default value is 1e-10.
Min_prob: Specify the minimum probability to use for observations without enough data. The default value is 0.001.
Eps_prob: Specify the threshold for standard deviation. If this threshold is not met, the min_sdev value is used. The default value is 1e-10.
Max_confusion_matrix_size: Specify the maximum size (in number of classes) for confusion matrices to be printed in the Logs.
Max_hit_ratio_k: Specify the maximum number (top K) of predictions to use for hit ratio computation. Applicable to multi-class only. To disable, enter 0.
Interpreting a Naïve Bayes Model
The output from Naïve Bayes is a list of tables containing the a-priori and conditional probabilities of each class of the response. The a-priori probability is the estimated probability of a particular class before observing any of the predictors. Each conditional probability table corresponds to a predictor column. The row headers are the classes of the response and the column headers are the classes of the predictor. Thus, in the table below, the probability of survival (y) given a person is male (x) is 0.91543624.
Sex
Survived Male Female
No 0.91543624 0.08456376
Yes 0.51617440 0.48382560
When the predictor is numeric, Naïve Bayes assumes it is sampled from a Gaussian distribution given the class of the response. The first column contains the mean and the second column contains the standard deviation of the distribution.
By default, the following output displays:
- Output (model category, model summary, scoring history, training metrics, validation metrics)
- Y-Levels (levels of the response column)
- P-conditionals
FAQ
How does the algorithm handle missing values during training?
All rows with one or more missing values (either in the predictors or the response) will be skipped during model building.
How does the algorithm handle missing values during testing?
If a predictor is missing, it will be skipped when taking the product of conditional probabilities in calculating the joint probability conditional on the response.
What happens if the response domain is different in the training and test datasets?
The response column in the test dataset is not used during scoring, so any response categories absent in the training data will not be predicted.
What happens during prediction if the new sample has categorical levels not seen in training?
The conditional probability of that predictor level will be set according to the Laplace smoothing factor. If Laplace smoothing is disabled (set to zero), the joint probability will be zero. See pgs. 13-14 of Andrew Ng’s “Generative learning algorithms” in the References section for mathematical details.
Does it matter if the data is sorted?
No.
Should data be shuffled before training?
This does not affect model building.
How does the algorithm handle highly imbalanced data in a response column?
Unbalanced data will not affect the model. However, if one response category has very few observations compared to the total, the conditional probability may be very low. A cutoff (
eps_prob) and minimum value (min_prob) are available for the user to set a floor on the calculated probability.
What if there are a large number of columns?
More memory will be allocated on each node to store the joint frequency counts and sums.
What if there are a large number of categorical factor levels?
More memory will be allocated on each node to store the joint frequency count of each categorical predictor level with the response’s level.
Naïve Bayes Algorithm
The algorithm is presented for the simplified binomial case without loss of generality.
Under the Naive Bayes assumption of independence, given a training set for a set of discrete valued features X \({(X^{(i)},\ y^{(i)};\ i=1,...m)}\)
The joint likelihood of the data can be expressed as:
\(\mathcal{L} \: (\phi(y),\: \phi_{i|y=1},\:\phi_{i|y=0})=\Pi_{i=1}^{m} p(X^{(i)},\: y^{(i)})\)
The model can be parameterized by:
\(\phi_{i|y=0}=\ p(x_{i}=1|\ y=0);\: \phi_{i|y=1}=\ p(x_{i}=1|y=1);\: \phi(y)\)
Where \(\phi_{i|y=0}=\ p(x_{i}=1|\ y=0)\) can be thought of as the fraction of the observed instances where feature \(x_{i}\) is observed, and the outcome is \(y=0, \phi_{i|y=1}=p(x_{i}=1|\ y=1)\) is the fraction of the observed instances where feature \(x_{i}\) is observed, and the outcome is \(y=1\), and so on.
The objective of the algorithm is to maximize with respect to \(\phi_{i|y=0}, \ \phi_{i|y=1},\ and \ \phi(y)\)
Where the maximum likelihood estimates are:
\(\phi_{j|y=1}= \frac{\Sigma_{i}^m 1(x_{j}^{(i)}=1 \ \bigcap y^{i} = 1)}{\Sigma_{i=1}^{m}(y^{(i)}=1}\)
\(\phi_{j|y=0}= \frac{\Sigma_{i}^m 1(x_{j}^{(i)}=1 \ \bigcap y^{i} = 0)}{\Sigma_{i=1}^{m}(y^{(i)}=0}\)
\(\phi(y)= \frac{(y^{i} = 1)}{m}\)
Once all parameters \(\phi_{j|y}\) are fitted, the model can be used to predict new examples with features \(X_{(i^*)}\).
This is carried out by calculating:
\(p(y=1|x)=\frac{\Pi p(x_i|y=1) p(y=1)}{\Pi p(x_i|y=1)p(y=1) \: +\: \Pi p(x_i|y=0)p(y=0)}\)
\(p(y=0|x)=\frac{\Pi p(x_i|y=0) p(y=0)}{\Pi p(x_i|y=1)p(y=1) \: +\: \Pi p(x_i|y=0)p(y=0)}\)
and predicting the class with the highest probability.
It is possible that prediction sets contain features not originally seen in the training set. If this occurs, the maximum likelihood estimates for these features predict a probability of 0 for all cases of y.
Laplace smoothing allows a model to predict on out of training data features by adjusting the maximum likelihood estimates to be:
\(\phi_{j|y=1}= \frac{\Sigma_{i}^m 1(x_{j}^{(i)}=1 \ \bigcap y^{i} = 1) \: + \: 1}{\Sigma_{i=1}^{m}(y^{(i)}=1 \: + \: 2}\)
\(\phi_{j|y=0}= \frac{\Sigma_{i}^m 1(x_{j}^{(i)}=1 \ \bigcap y^{i} = 0) \: + \: 1}{\Sigma_{i=1}^{m}(y^{(i)}=0 \: + \: 2}\)
Note that in the general case where y takes on k values, there are k+1 modified parameter estimates, and they are added in when the denominator is k (rather than two, as shown in the two-level classifier shown here.)
Laplace smoothing should be used with care; it is generally intended to allow for predictions in rare events. As prediction data becomes increasingly distinct from training data, train new models when possible to account for a broader set of possible X values.
References
Ng, Andrew. “Generative Learning algorithms.” (2008).
PCA
PCA is currently in progress in H2O. Once implementation of this algorithm is complete, this section of the document will be updated.
GBM
Introduction
Gradient Boosted Regression and Gradient Boosted Classification are forward learning ensemble methods. The guiding heuristic is that good predictive results can be obtained through increasingly refined approximations. H2O’s GBM sequentially builds regression trees on all the features of the dataset in a fully distributed way - each tree is built in parallel.
Defining a GBM Model
Destination_key: (Optional) Enter a custom name for the model to use as a reference. By default, H2O automatically generates a destination key.
Training_frame: (Required) Select the dataset used to build the model. NOTE: If you click the Build a model button from the
Parsecell, the training frame is entered automatically.Validation_frame: (Optional) Select the dataset used to evaluate the accuracy of the model.
Ignored_columns: (Optional) Click the plus sign next to a column name to add it to the list of columns excluded from the model. To add all columns, click the Add all button. To remove a column from the list of ignored columns, click the X next to the column name. To remove all columns from the list of ignored columns, click the Clear all button.
Ignore_const_cols: Check this checkbox to ignore constant training columns, since no information can be gained from them. This option is selected by default.
Score_each_iteration: (Optional) Check this checkbox to score during each iteration of the model training.
Response_column: (Required) Select the column to use as the independent variable.
Ntrees: Specify the number of trees. The default value is 50.
Max_depth: Specify the maximum tree depth. The default value is 5.
Min_rows: Specify the minimum number of observations for a leaf (
nodesizein R). The default value is 10.Nbins: Specify the number of bins for the histogram. The default value is 20.
Learn_rate: Specify the learning rate. The range is 0.0 to 1.0 and the default is 0.1.
Distribution: Select the loss function. The options are auto, bernoulli, multinomial, or gaussian and the default is auto.
Balance_classes: Oversample the minority classes to balance the class distribution. This option is not selected by default. This option is only applicable for classification. Majority classes can be undersampled to satisfy the Max_after_balance_size parameter.
Max_confusion_matrix_size: Specify the maximum size (in number of classes) for confusion matrices to be printed in the Logs.
Max_hit_ratio_k: Specify the maximum number (top K) of predictions to use for hit ratio computation. Applicable to multi-class only. To disable, enter 0.
Class_sampling_factors: Specify the per-class (in lexicographical order) over/under-sampling ratios. By default, these ratios are automatically computed during training to obtain the class balance. There is no default value.
Seed: Specify the random number generator (RNG) seed for algorithm components dependent on randomization. The seed is consistent for each H2O instance so that you can create models with the same starting conditions in alternative configurations.
Interpreting a GBM Model
The output for GBM includes the following:
- Model parameters (hidden)
- A graph of the scoring history (training MSE vs number of trees)
- A graph of the variable importances
- Output (model category, validation metrics, initf)
- Model summary (number of trees, min. depth, max. depth, mean depth, min. leaves, max. leaves, mean leaves)
- Scoring history in tabular format
- Training metrics (model name, model checksum name, frame name, description, model category, duration in ms, scoring time, predictions, MSE, R2)
- Variable importances in tabular format
FAQ
How does the algorithm handle missing values during training?
Missing values do not alter the tree building in any way (i.e., they are not counted as a point when computing means or errors). Rows containing missing values do affect tree building, but the missing values don’t change the split-point of the column they are in.
How does the algorithm handle missing values during testing?
During scoring, missing values “always go left” at any decision point in a tree. Due to dynamic binning in GBM, a row with a missing value typically ends up in the “leftmost bin” - with other outliers.
What happens if the response has missing values?
No errors will occur, but nothing will be learned from rows containing missing the response.
Does it matter if the data is sorted?
No.
Should data be shuffled before training?
No.
How does the algorithm handle highly imbalanced data in a response column?
You can specify
balance_classes,class_sampling_factorsandmax_after_balance_sizeto control over/under-sampling.What if there are a large number of columns?
DRF models are best for datasets with fewer than a few thousand columns.
What if there are a large number of categorical factor levels?
Large number of categoricals are handled very efficiently - there is never any one-hot encoding.
GBM Algorithm
H2O’s Gradient Boosting Algorithms follow the algorithm specified by Hastie et al (2001):
Initialize \(f_{k0} = 0,\: k=1,2,…,K\)
For \(m=1\) to \(M:\)
(a) Set \(p_{k}(x)=\frac{e^{f_{k}(x)}}{\sum_{l=1}^{K}e^{f_{l}(x)}},\:k=1,2,…,K\)
(b) For \(k=1\) to \(K\):
i. Compute \(r_{ikm}=y_{ik}-p_{k}(x_{i}),\:i=1,2,…,N.\) ii. Fit a regression tree to the targets \(r_{ikm},\:i=1,2,…,N\), giving terminal regions \(R_{jim},\:j=1,2,…,J_{m}.\) \(iii. Compute\) \(\gamma_{jkm}=\frac{K-1}{K}\:\frac{\sum_{x_{i}\in R_{jkm}}(r_{ikm})}{\sum_{x_{i}\in R_{jkm}}|r_{ikm}|(1-|r_{ikm})},\:j=1,2,…,J_{m}.\) \(\:iv.\:Update\:f_{km}(x)=f_{k,m-1}(x)+\sum_{j=1}^{J_{m}}\gamma_{jkm}I(x\in\:R_{jkm}).\)
Output \(\:\hat{f_{k}}(x)=f_{kM}(x),\:k=1,2,…,K.\)
References
Dietterich, Thomas G, and Eun Bae Kong. “Machine Learning Bias, Statistical Bias, and Statistical Variance of Decision Tree Algorithms.” ML-95 255 (1995).
Elith, Jane, John R Leathwick, and Trevor Hastie. “A Working Guide to Boosted Regression Trees.” Journal of Animal Ecology 77.4 (2008): 802-813
Friedman, Jerome H. “Greedy Function Approximation: A Gradient Boosting Machine.” Annals of Statistics (2001): 1189-1232.
Friedman, Jerome, Trevor Hastie, Saharon Rosset, Robert Tibshirani, and Ji Zhu. “Discussion of Boosting Papers.” Ann. Statist 32 (2004): 102-107
Deep Learning
- Introduction
- Defining a Deep Learning Model
- Interpreting a Deep Learning Model
- FAQ
- Deep Learning Algorithm
- References
Introduction
H2O’s Deep Learning is based on a multi-layer feed-forward artificial neural network that is trained with stochastic gradient descent using back-propagation. The network can contain a large number of hidden layers consisting of neurons with tanh, rectifier and maxout activation functions. Advanced features such as adaptive learning rate, rate annealing, momentum training, dropout, L1 or L2 regularization, checkpointing and grid search enable high predictive accuracy. Each compute node trains a copy of the global model parameters on its local data with multi-threading (asynchronously), and contributes periodically to the global model via model averaging across the network.
Defining a Deep Learning Model
H2O Deep Learning models have many input parameters, many of which are only accessible via the expert mode. For most cases, use the default values. Please read the following instructions before building extensive Deep Learning models. The application of grid search and successive continuation of winning models via checkpoint restart is highly recommended, as model performance can vary greatly.
Destination_key: (Optional) Enter a custom name for the model to use as a reference. By default, H2O automatically generates a destination key.
Training_frame: (Required) Select the dataset used to build the model. NOTE: If you click the Build a model button from the
Parsecell, the training frame is entered automatically.Validation_frame: (Optional) Select the dataset used to evaluate the accuracy of the model.
Ignored_columns: (Optional) Click the plus sign next to a column name to add it to the list of columns excluded from the model. To add all columns, click the Add all button. To remove a column from the list of ignored columns, click the X next to the column name. To remove all columns from the list of ignored columns, click the Clear all button.
Ignore_const_cols: Check this checkbox to ignore constant training columns, since no information can be gained from them. This option is selected by default.
Response_column: Select the column to use as the independent variable.
Activation: Select the activation function (Tahn, Tahn with dropout, Rectifier, Rectifier with dropout, Maxout, Maxout with dropout). The default option is Rectifier.
Hidden: Specify the hidden layer sizes (e.g., 100,100). The default value is 200,200.
Epochs: Specify the number of times to iterate (stream) the dataset. The value can be a fraction. The default value for DL is 10.
Variable_importances: Check this checkbox to compute variable importance. This option is not selected by default.
Balance_classes: Oversample the minority classes to balance the class distribution. This option is not selected by default. This option is only applicable for classification. Majority classes can be undersampled to satisfy the Max_after_balance_size parameter.
Max_confusion_matrix_size: Specify the maximum size (in number of classes) for confusion matrices to be printed in the Logs.
Max_hit_ratio_k: Specify the maximum number (top K) of predictions to use for hit ratio computation. Applicable to multi-class only. To disable, enter 0.
Checkpoint: Enter a model key associated with a previously-trained Deep Learning model. Use this option to build a new model as a continuation of a previously-generated model (e.g., by a grid search).
Use_all_factor_levels: Check this checkbox to use all factor levels in the possible set of predictors; if you enable this option, sufficient regularization is required. By default, the first factor level is skipped. For Deep Learning models, this option is useful for determining variable importances and is automatically enabled if the autoencoder is selected.
Train_samples_per_iteration: Specify the number of global training samples per MapReduce iteration. To specify one epoch, enter 0. To specify all available data (e.g., replicated training data), enter -1. To use the automatic values, enter -2. The default is -2.
Adaptive_rate: Check this checkbox to enable the adaptive learning rate (ADADELTA). This option is selected by default.
Input_dropout_ratio: Specify the input layer dropout ratio to improve generalization. Suggested values are 0.1 or 0.2. The default value is 0.
L1: Specify the L1 regularization to add stability and improve generalization; sets the value of many weights to 0. The default value is 0.
L2: Specify the L2 regularization to add stability and improve generalization; sets the value of many weights to smaller values. The default value is 0.
Loss: Select the loss function. The options are automatic, mean square, cross-entropy, Huber, or Absolute and the default value is automatic.
Score_interval: Specify the shortest time interval (in seconds) to wait between model scoring. The default value is 5.0.
Score_training_samples: Specify the number of training set samples for scoring. To use all training samples, enter 0. The default value is 10000.
Score_duty_cycle: Specify the maximum duty cycle fraction for scoring. A lower value results in more training and a higher value results in more scoring. The default value is 0.1.
Autoencoder: Check this checkbox to enable the Deep Learning autoencoder. This option is not selected by default. Note: This option requires MeanSquare as the loss function.
Class_sampling_factors: Specify the per-class (in lexicographical order) over/under-sampling ratios. By default, these ratios are automatically computed during training to obtain the class balance. There is no default value.
Overwrite_with_best_model: Check this checkbox to overwrite the final model with the best model found during training. This option is selected by default.
Target_ratio_comm_to_comp: Specify the target ratio of communication overhead to computation. This option is only enabled for multi-node operation and if train_samples_per_iteration equals -2 (auto-tuning). The default value is 0.02.
Seed: Specify the random number generator (RNG) seed for algorithm components dependent on randomization. The seed is consistent for each H2O instance so that you can create models with the same starting conditions in alternative configurations.
Rho: Specify the adaptive learning rate time decay factor. The default value is 0.99.
Epsilon: Specify the adaptive learning rate time smoothing factor to avoid dividing by zero. The default value is 1e-8.
Max_W2: Specify the constraint for the squared sum of the incoming weights per unit (e.g., for Rectifier). The default value is infinity.
Initial_weight_distribution: Select the initial weight distribution (Uniform Adaptive, Uniform, or Normal). The default is Uniform Adaptive.
Regression_stop: Specify the stopping criterion for regression error (MSE) on the training data. To disable this option, enter -1. The default value is 1.0E-6.
Diagnostics: Check this checkbox to compute the variable importances for input features (using the Gedeon method). For large networks, selecting this option can reduce speed. This option is selected by default.
Fast_mode: Check this checkbox to enable fast mode, a minor approximation in back-propagation. This option is selected by default.
Ignore_const_cols: Check this checkbox to ignore constant training columns, since no information can be gained from them. This option is selected by default.
Force_load_balance: Check this checkbox to force extra load balancing to increase training speed for small datasets and use all cores. This option is selected by default.
Single_node_mode: Check this checkbox to force H2O to run on a single node for fine-tuning of model parameters. This option is not selected by default.
Shuffle_training_data: Check this checkbox to shuffle the training data. This option is recommended if the training data is replicated and the value of train_samples_per_iteration is close to the number of nodes times the number of rows. This option is not selected by default.
Missing_values_handling: Select how to handle missing values (skip or mean imputation). The default value is mean imputation.
Quiet_mode: Check this checkbox to display less output in the standard output. This option is not selected by default.
Sparse: Check this checkbox to use sparse data handling. This option is not selected by default.
Col_major: Check this checkbox to use a column major weight matrix for the input layer. This option can speed up forward propagation but may reduce the speed of backpropagation. This option is not selected by default.
Average_activation: Specify the average activation for the sparse autoencoder. The default value is 0.0.
Sparsity_beta: Specify the sparsity regularization. The default value is 0.0.
Max_categorical_features: Specify the maximum number of categorical features enforced via hashing.
Reproducible: To force reproducibility on small data, check this checkbox. If this option is enabled, the model takes more time to generate, since it uses only one thread.
Export_weights_and_biases: To export the neural network weights and biases as H2O frames, check this checkbox.
Interpreting a Deep Learning Model
To view the results, click the View button. The output for the Deep Learning model includes the following information for both the training and testing sets:
- Model parameters (hidden)
- A chart of the variable importances
- A graph of the scoring history (training MSE and validation MSE vs epochs)
- Output (model category, weights, biases)
- Status of neuron layers (layer number, units, type, dropout, L1, L2, mean rate, rate RMS, momentum, mean weight, weight RMS, mean bias, bias RMS)
- Scoring history in tabular format
- Training metrics (model name, model checksum name, frame name, frame checksum name, description, model category, duration in ms, scoring time, predictions, MSE, R2, logloss)
- Top-K Hit Ratios (for multi-class classification)
- Confusion matrix (for classification)
FAQ
How does the algorithm handle missing values during training?
User-specifiable treatment of missing values via
missing_values_handling. Specify either the skip or mean-impute option.How does the algorithm handle missing values during testing?
Missing values in the test set will be mean-imputed during scoring.
What happens if the response has missing values?
No errors will occur, but nothing will be learned from rows containing missing the response.
Does it matter if the data is sorted?
Yes, since the training set is processed in order. Depending whether
train_samples_per_iterationis enabled, some rows will be skipped. Ifshuffle_training_datais enabled, then each thread that is processing a small subset of rows will process rows randomly, but it is not a global shuffle.Should data be shuffled before training?
Yes, the data should be shuffled before training, especially if the dataset is sorted.
How does the algorithm handle highly imbalanced data in a response column?
Specify
balance_classes,class_sampling_factorsandmax_after_balance_sizeto control over/under-sampling.What if there are a large number of columns?
The input neuron layer’s size is scaled to the number of input features, so as the number of columns increases, the model complexity increases as well.
What if there are a large number of categorical factor levels?
This is something to look out for. Say you have three columns: zip code (70k levels), height, and income. The resulting number of internally one-hot encoded features will be 70,002 and only 3 of them will be activated (non-zero). If the first hidden layer has 200 neurons, then the resulting weight matrix will be of size 70,002 x 200, which can take a long time to train and converge. In this case, we recommend either reducing the number of categorical factor levels upfront (e.g., using h2o.interaction() from R), or specifying max_categorical_features to use feature hashing to reduce the dimensionality.
Deep Learning Algorithm
For more information about how the Deep Learning algorithm works, refer to the Deep Learning booklet.
References
Candel, Arno and Parmar, Viraj. “Deep Learning with H2O.” H2O.ai, Inc. (2015).
Candel, Arno. “The Definitive Performance Tuning Guide for H2O Deep Learning.” H2O.ai, Inc. (2015).
YARN Best Practices
- Using H2O with YARN
- Configuring YARN
- Limiting CPU Usage
- Specifying Queues
- Specifying Output Directories
- Customizing YARN
- Accessing Logs
YARN (Yet Another Resource Manager) is a resource management framework. H2O can be launched as an application on YARN. If you want to run H2O on Hadoop, essentially, you are running H2O on YARN. If you are not currently using YARN to manage your cluster resources, we strongly recommend it.
Using H2O with YARN
When you launch H2O on Hadoop using the hadoop jar command, YARN allocates the necessary resources to launch the requested number of nodes. H2O launches as a MapReduce (V2) task, where each mapper is an H2O node of the specified size.
Occasionally, YARN may reject a job request. This usually occurs because either there is not enough memory to launch the job or because of an incorrect configuration.
If YARN rejects the job request, try launching the job with less memory to see if that is the cause of the failure. Specify smaller values (we recommend 1) for -mapperXmx and -nodes to confirm that H2O can launch successfully:
hadoop jar h2odriver.jar -nodes 1 -mapperXmx 1g -output hdfsOutputDirName
To resolve configuration issues, adjust the maximum memory that YARN will allow when launching each mapper. If the cluster manager settings are configured for the default maximum memory size but the memory required for the request exceeds that amount, YARN will not launch and H2O will time out. If you are using the default configuration, change the configuration settings in your cluster manager to specify memory allocation when launching mapper tasks. To calculate the amount of memory required for a successful launch, use the following formula:
YARN container size (
mapreduce.map.memory.mb) =-mapperXmxvalue + (-mapperXmx*-extramempercent[default is 10%])
The mapreduce.map.memory.mb value must be less than the YARN memory configuration values for the launch to succeed.
Configuring YARN
For Cloudera, configure the settings in Cloudera Manager. Depending on how the cluster is configured, you may need to change the settings for more than one role group.
Click Configuration and enter the following search term in quotes: yarn.nodemanager.resource.memory-mb.
Enter the amount of memory (in GB) to allocate in the Value field. If more than one group is listed, change the values for all listed groups.
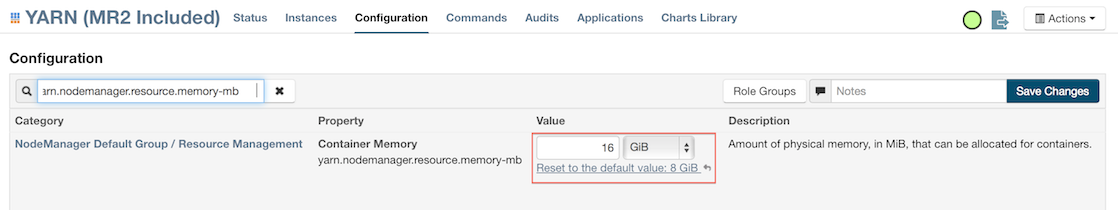
Click the Save Changes button in the upper-right corner.
- Enter the following search term in quotes: yarn.scheduler.maximum-allocation-mb
Change the value, click the Save Changes button in the upper-right corner, and redeploy.
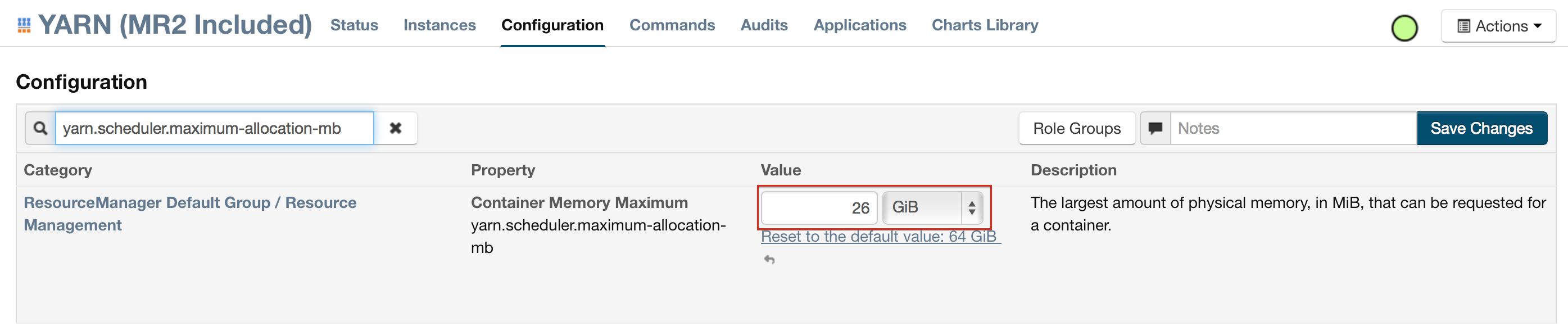
For Hortonworks, configure the settings in Ambari.
- Select YARN, then click the Configs tab.
- Select the group.
In the Node Manager section, enter the amount of memory (in MB) to allocate in the yarn.nodemanager.resource.memory-mb entry field.
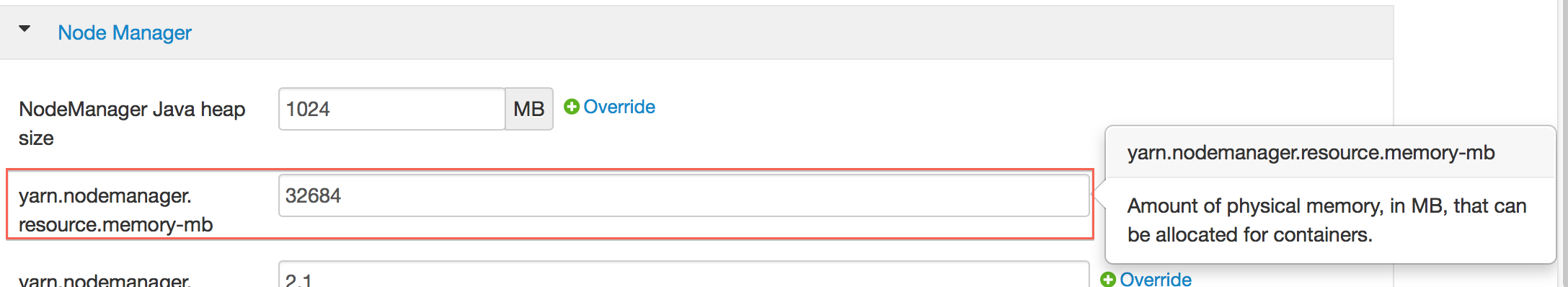
In the Scheduler section, enter the amount of memory (in MB)to allocate in the yarn.scheduler.maximum-allocation-mb entry field.
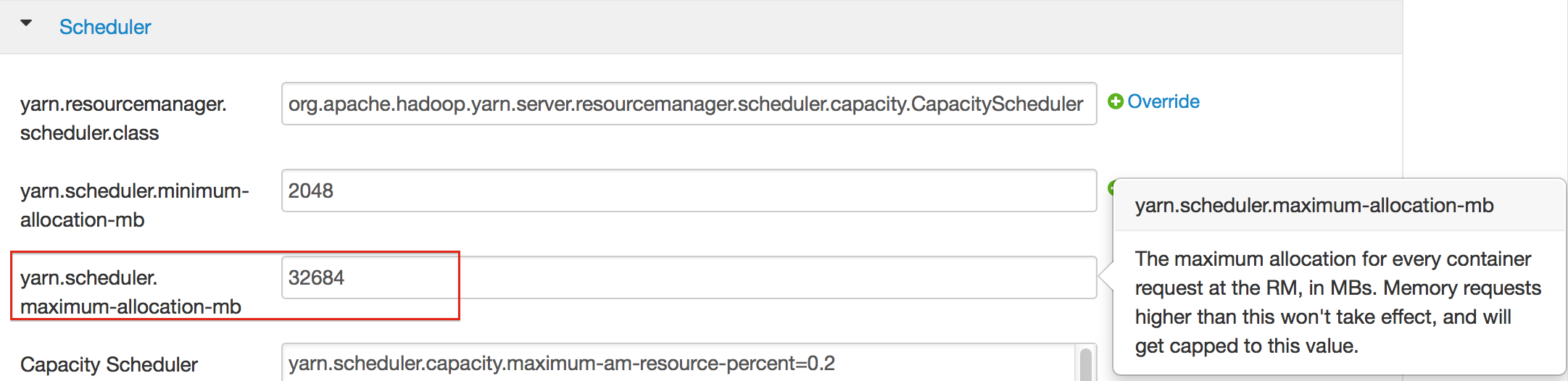
Click the Save button at the bottom of the page and redeploy the cluster.
For MapR:
- Edit the yarn-site.xml file for the node running the ResourceManager.
- Change the values for the
yarn.nodemanager.resource.memory-mbandyarn.scheduler.maximum-allocation-mbproperties. - Restart the ResourceManager and redeploy the cluster.
To verify the values were changed, check the values for the following properties:
- <name>yarn.nodemanager.resource.memory-mb</name>
- <name>yarn.scheduler.maximum-allocation-mb</name>
Limiting CPU Usage
To limit the number of CPUs used by H2O, use the -nthreads option and specify the maximum number of CPUs for a single container to use. The following example limits the number of CPUs to two:
java -jar h2o.jar -nthreads 2
The default is 4*the number of CPUs.
Specifying Queues
If you do not specify a queue when launching H2O, H2O jobs are submitted to the default queue. Jobs submitted to the default queue have a lower priority than jobs submitted to a specific queue.
To specify a queue with Hadoop, enter -Dmapreduce.job.queuename=<queue name>
(where <queue name> is the name of the queue) when launching Hadoop.
For example,
hadoop jar h2odriver.jar -Dmapreduce.job.queuename=default -nodes 1 -mapperXmx 1g -output hdfsOutputDirName.
Specifying Output Directories
To prevent overwriting multiple users’ files, each job must have a unique output directory name. Change the -output hdfsOutputDir argument (where hdfsOutputDir is the name of the directory.
Alternatively, you can delete the directory (manually or by using a script) instead of creating a unique directory each time you launch H2O.
Customizing YARN
Most of the configurable YARN variables are stored in yarn-site.xml. To prevent settings from being overridden, you can mark a config as “final.” If you change any values in yarn-site.xml, you must restart YARN to confirm the changes.
Accessing Logs
To learn how to access logs in YARN, refer to Downloading Logs.
Downloading Logs
Accessing Logs
Depending on whether you are using Hadoop with H2O and whether the job is currently running, there are different ways of obtaining the logs for H2O.
Copy and email the logs to support@h2o.ai or submit them to h2ostream@googlegroups.com with a brief description of your Hadoop environment, including the Hadoop distribution and version.
Without Running Jobs
- If you are using Hadoop and the job is not running, view the logs by using the
yarn logs -applicationIdcommand. When you start an H2O instance, the complete command displays in the output:
jessica@mr-0x8:~/h2o-3.1.0.3008-cdh5.2$ hadoop jar h2odriver.jar -nodes 1 -mapperXmx 1g -output hdfsOutputDirName
Determining driver host interface for mapper->driver callback...
[Possible callback IP address: 172.16.2.178]
[Possible callback IP address: 127.0.0.1]
Using mapper->driver callback IP address and port: 172.16.2.178:52030
(You can override these with -driverif and -driverport.)
Memory Settings:
mapreduce.map.java.opts: -Xms1g -Xmx1g -Dlog4j.defaultInitOverride=true
Extra memory percent: 10
mapreduce.map.memory.mb: 1126
15/05/06 17:11:50 INFO client.RMProxy: Connecting to ResourceManager at mr-0x10.0xdata.loc/172.16.2.180:8032
15/05/06 17:11:52 INFO mapreduce.JobSubmitter: number of splits:1
15/05/06 17:11:52 INFO mapreduce.JobSubmitter: Submitting tokens for job: job_1430127035640_0075
15/05/06 17:11:52 INFO impl.YarnClientImpl: Submitted application application_1430127035640_0075
15/05/06 17:11:52 INFO mapreduce.Job: The url to track the job: http://mr-0x10.0xdata.loc:8088/proxy/application_1430127035640_0075/
Job name 'H2O_29570' submitted
JobTracker job ID is 'job_1430127035640_0075'
For YARN users, logs command is 'yarn logs -applicationId application_1430127035640_0075'
Waiting for H2O cluster to come up...
In the above example, the command is specified in the next to last line (For YARN users, logs command is...). The command is unique for each instance. In Terminal, enter yarn logs -applicationId application_<UniqueID> to view the logs (where <UniqueID> is the number specified in the next to last line of the output that displayed when you created the cluster).
Use YARN to obtain the stdout and stderr logs that are used for troubleshooting. To learn how to access YARN based on management software, version, and job status, see Accessing YARN.
Click the Applications link to view all jobs, then click the History link for the job.
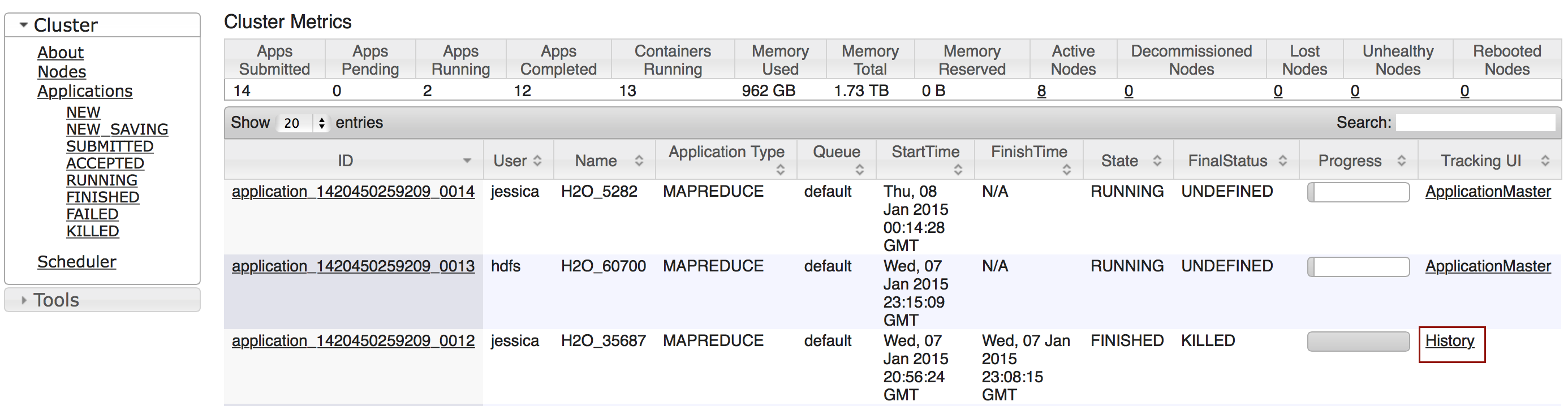
Click the logs link.
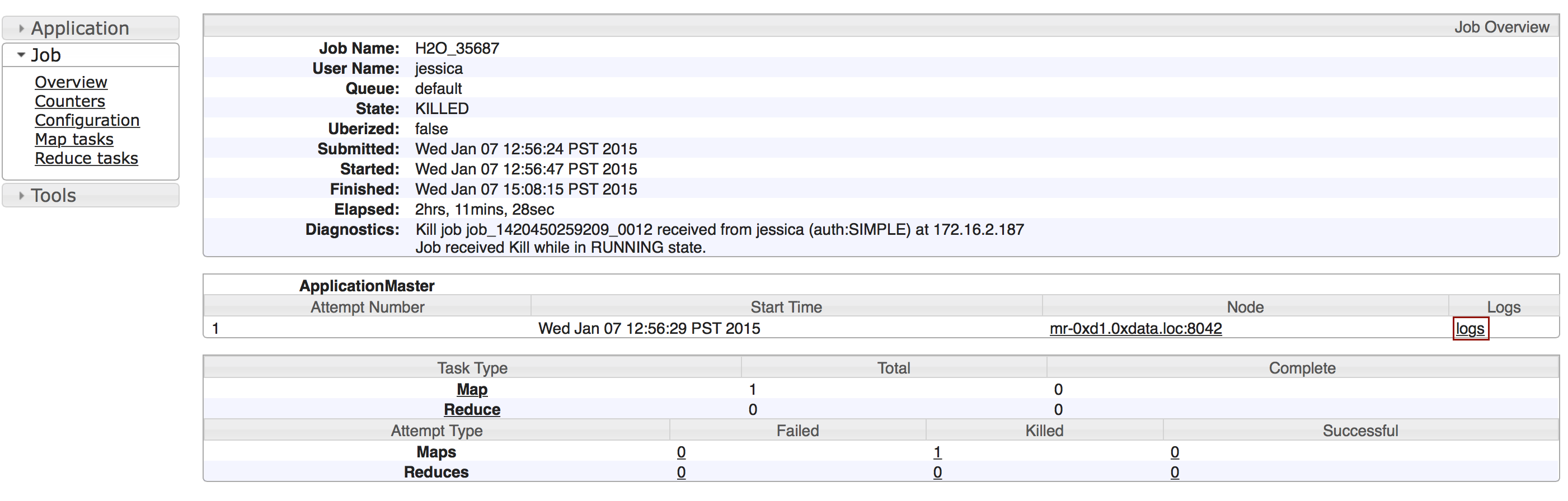
Copy the information that displays and send it in an email to support@h2o.ai.
With Running Jobs
If you are using Hadoop and the job is still running:
Use YARN to obtain the
stdoutandstderrlogs that are used for troubleshooting. To learn how to access YARN based on management software, version, and job status, see Accessing YARN.Click the Applications link to view all jobs, then click the ApplicationMaster link for the job.
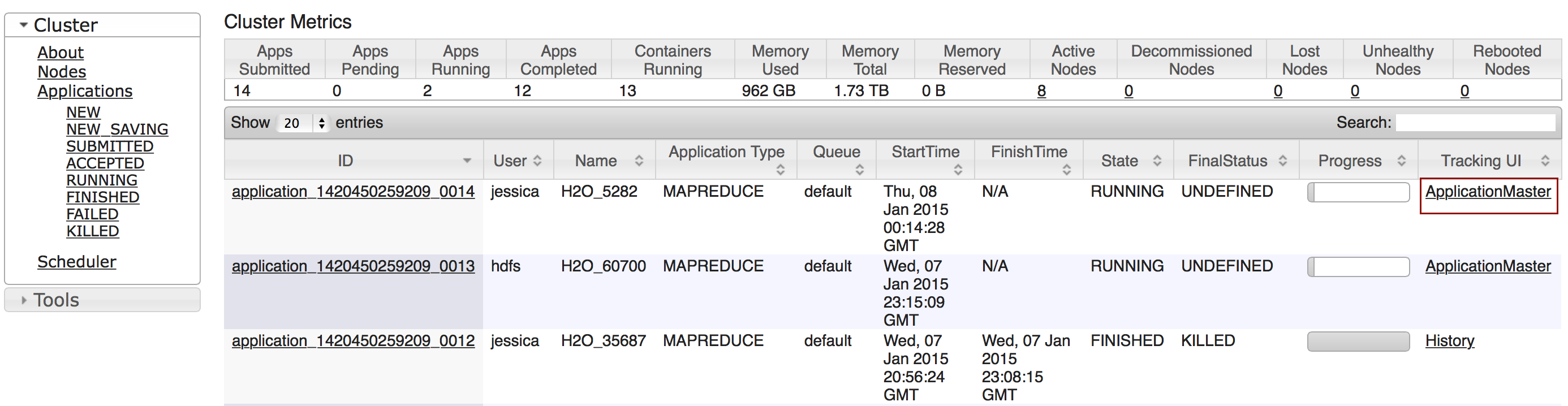
Select the job from the list of active jobs.
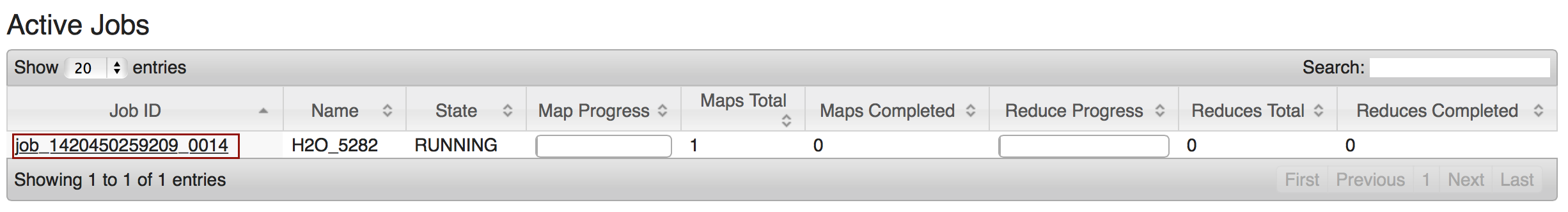
Click the logs link.
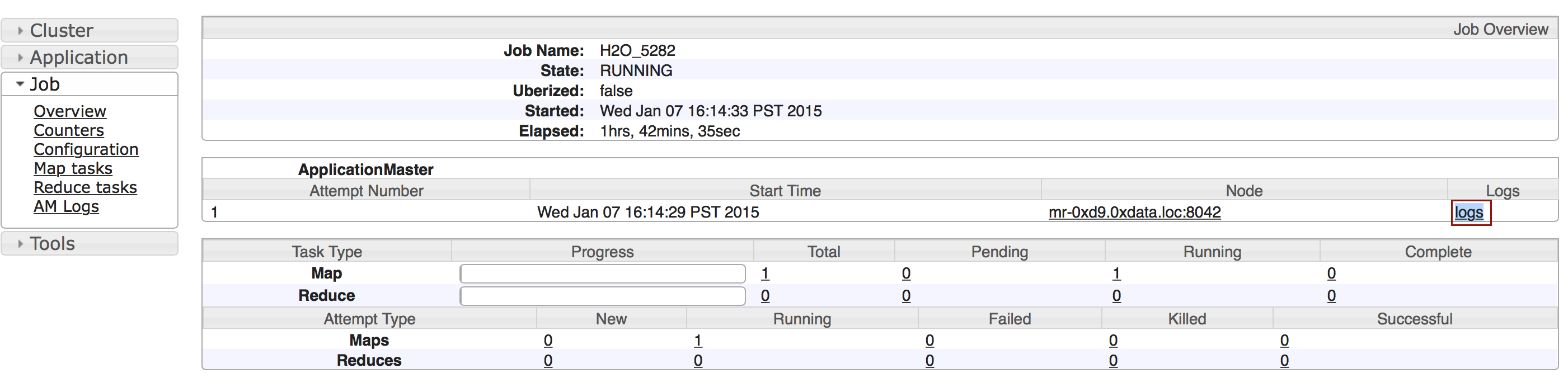
Send the contents of the displayed files to support@h2o.ai.

Go to the H2O web UI and select Admin > View Log. To filter the results select a node or log file type from the drop-down menus. To download the logs, click the Download Logs button.
When you view the log, the output displays the location of log directory after
Log dir:(as shown in the last line in the following example):
05-06 17:12:15.610 172.16.2.179:54321 26336 main INFO: ----- H2O started -----
05-06 17:12:15.731 172.16.2.179:54321 26336 main INFO: Build git branch: master
05-06 17:12:15.731 172.16.2.179:54321 26336 main INFO: Build git hash: 41d039196088df081ad77610d3e2d6550868f11b
05-06 17:12:15.731 172.16.2.179:54321 26336 main INFO: Build git describe: jenkins-master-1187
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Build project version: 0.3.0.1187
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Built by: 'jenkins'
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Built on: '2015-05-05 23:31:12'
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Java availableProcessors: 8
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Java heap totalMemory: 982.0 MB
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Java heap maxMemory: 982.0 MB
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Java version: Java 1.7.0_80 (from Oracle Corporation)
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: OS version: Linux 3.13.0-51-generic (amd64)
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Machine physical memory: 31.30 GB
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: X-h2o-cluster-id: 1430957535344
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Possible IP Address: virbr0 (virbr0), 192.168.122.1
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Possible IP Address: br0 (br0), 172.16.2.179
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Possible IP Address: lo (lo), 127.0.0.1
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Multiple local IPs detected:
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: /192.168.122.1 /172.16.2.179
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Attempting to determine correct address...
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Using /172.16.2.179
05-06 17:12:15.734 172.16.2.179:54321 26336 main INFO: Internal communication uses port: 54322
05-06 17:12:15.734 172.16.2.179:54321 26336 main INFO: Listening for HTTP and REST traffic on http://172.16.2.179:54321/
05-06 17:12:15.744 172.16.2.179:54321 26336 main INFO: H2O cloud name: 'H2O_29570' on /172.16.2.179:54321, discovery address /237.61.246.13:60733
05-06 17:12:15.744 172.16.2.179:54321 26336 main INFO: If you have trouble connecting, try SSH tunneling from your local machine (e.g., via port 55555):
05-06 17:12:15.744 172.16.2.179:54321 26336 main INFO: 1. Open a terminal and run 'ssh -L 55555:localhost:54321 yarn@172.16.2.179'
05-06 17:12:15.744 172.16.2.179:54321 26336 main INFO: 2. Point your browser to http://localhost:55555
05-06 17:12:15.979 172.16.2.179:54321 26336 main INFO: Log dir: '/home2/yarn/nm/usercache/jessica/appcache/application_1430127035640_0075/h2ologs'
- In Terminal, enter
cd /tmp/h2o-<UserName>/h2ologs(where<UserName>is your computer user name), then enterls -lto view a list of the log files. Thehttpdlog contains the request/response status of all REST API transactions. The rest of the logs use the formath2o_\<IPaddress>\_<Port>-<LogLevel>-<LogLevelName>.log, where<IPaddress>is the bind address of the H2O instance,<Port>is the port number,<LogLevel>is the numerical log level (1-6, with 6 as the highest severity level), and<LogLevelName>is the name of the log level (trace, debug, info, warn, error, or fatal).
- Download the logs using R. In R, enter the command
h2o.downloadAllLogs(client = localH2O,filename = "logs.zip")(whereclientis the H2O cluster andfilenameis the specified filename for the logs).
Accessing YARN
Methods for accessing YARN vary depending on the default management software and version, as well as job status.
Cloudera 5 & 5.2
In Cloudera Manager, click the YARN link in the cluster section.
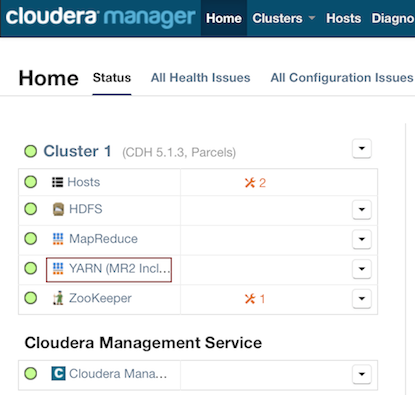
In the Quick Links section, select ResourceManager Web UI if the job is running or select HistoryServer Web UI if the job is not running.
Ambari
From the Ambari Dashboard, select YARN.
From the Quick Links drop-down menu, select ResourceManager UI.
For Non-Hadoop Users
Without Current Jobs
If you are not using Hadoop and the job is not running:
- In Terminal, enter
cd /tmp/h2o-<UserName>/h2ologs(where<UserName>is your computer user name), then enterls -lto view a list of the log files. Thehttpdlog contains the request/response status of all REST API transactions. The rest of the logs use the formath2o_\<IPaddress>\_<Port>-<LogLevel>-<LogLevelName>.log, where<IPaddress>is the bind address of the H2O instance,<Port>is the port number,<LogLevel>is the numerical log level (1-6, with 6 as the highest severity level), and<LogLevelName>is the name of the log level (trace, debug, info, warn, error, or fatal).
With Current Jobs
If you are not using Hadoop and the job is still running:
Go to the H2O web UI and select Admin > Inspect Log or go to http://localhost:54321/LogView.html.
To download the logs, click the Download Logs button.
When you view the log, the output displays the location of log directory after
Log dir:(as shown in the last line in the following example):
05-06 17:12:15.610 172.16.2.179:54321 26336 main INFO: ----- H2O started -----
05-06 17:12:15.731 172.16.2.179:54321 26336 main INFO: Build git branch: master
05-06 17:12:15.731 172.16.2.179:54321 26336 main INFO: Build git hash: 41d039196088df081ad77610d3e2d6550868f11b
05-06 17:12:15.731 172.16.2.179:54321 26336 main INFO: Build git describe: jenkins-master-1187
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Build project version: 0.3.0.1187
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Built by: 'jenkins'
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Built on: '2015-05-05 23:31:12'
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Java availableProcessors: 8
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Java heap totalMemory: 982.0 MB
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Java heap maxMemory: 982.0 MB
05-06 17:12:15.732 172.16.2.179:54321 26336 main INFO: Java version: Java 1.7.0_80 (from Oracle Corporation)
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: OS version: Linux 3.13.0-51-generic (amd64)
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Machine physical memory: 31.30 GB
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: X-h2o-cluster-id: 1430957535344
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Possible IP Address: virbr0 (virbr0), 192.168.122.1
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Possible IP Address: br0 (br0), 172.16.2.179
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Possible IP Address: lo (lo), 127.0.0.1
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Multiple local IPs detected:
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: /192.168.122.1 /172.16.2.179
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Attempting to determine correct address...
05-06 17:12:15.733 172.16.2.179:54321 26336 main INFO: Using /172.16.2.179
05-06 17:12:15.734 172.16.2.179:54321 26336 main INFO: Internal communication uses port: 54322
05-06 17:12:15.734 172.16.2.179:54321 26336 main INFO: Listening for HTTP and REST traffic on http://172.16.2.179:54321/
05-06 17:12:15.744 172.16.2.179:54321 26336 main INFO: H2O cloud name: 'H2O_29570' on /172.16.2.179:54321, discovery address /237.61.246.13:60733
05-06 17:12:15.744 172.16.2.179:54321 26336 main INFO: If you have trouble connecting, try SSH tunneling from your local machine (e.g., via port 55555):
05-06 17:12:15.744 172.16.2.179:54321 26336 main INFO: 1. Open a terminal and run 'ssh -L 55555:localhost:54321 yarn@172.16.2.179'
05-06 17:12:15.744 172.16.2.179:54321 26336 main INFO: 2. Point your browser to http://localhost:55555
05-06 17:12:15.979 172.16.2.179:54321 26336 main INFO: Log dir: '/home2/yarn/nm/usercache/jessica/appcache/application_1430127035640_0075/h2ologs'
- In Terminal, enter
cd /tmp/h2o-<UserName>/h2ologs(where<UserName>is your computer user name), then enterls -lto view a list of the log files. Thehttpdlog contains the request/response status of all REST API transactions. The rest of the logs use the formath2o_\<IPaddress>\_<Port>-<LogLevel>-<LogLevelName>.log, where<IPaddress>is the bind address of the H2O instance,<Port>is the port number,<LogLevel>is the numerical log level (1-6, with 6 as the highest severity level), and<LogLevelName>is the name of the log level (trace, debug, info, warn, error, or fatal).
To view the REST API logs from R:
In R, enter
h2o.startLogging(). The output displays the location of the REST API logs:> h2o.startLogging() Appending REST API transactions to log file /var/folders/ylcq5nhky53hjcl9wrqxt39kz80000gn/T//RtmpE7X8Yv/rest.logCopy the displayed file path. Enter
lessand paste the file path.- Press Enter. A time-stamped log of all REST API transactions displays.
------------------------------------------------------------
Time: 2015-01-06 15:46:11.083
GET http://172.16.2.20:54321/3/Cloud.json
postBody:
curlError: FALSE
curlErrorMessage:
httpStatusCode: 200
httpStatusMessage: OK
millis: 3
{"__meta":{"schema_version": 1,"schema_name":"CloudV1","schema_type":"Iced"},"version":"0.1.17.1009","cloud_name":...[truncated]}
-------------------------------------------------------------
- Download the logs using R. In R, enter the command
h2o.downloadAllLogs(client = localH2O,filename = "logs.zip")(whereclientis the H2O cluster andfilenameis the specified filename for the logs).
Migrating to H2O 3.0
We’re excited about the upcoming release of the latest and greatest version of H2O, and we hope you are too! H2O 3.0 has lots of improvements, including:
- Powerful Python APIs
- Flow, a brand-new intuitive web UI
- The ability to share, annotate, and modify workflows
- Versioned REST APIs with full metadata
- Spark integration using Sparkling Water
- Improved algorithm accuracy and speed
and much more! Overall, H2O has been retooled for better accuracy and performance and to provide additional functionality. If you’re a current user of H2O, we strongly encourage you to upgrade to the latest version to take advantage of the latest features and capabilities.
Please be aware that H2O 3.0 will supersede all previous versions of H2O as the primary version as of May 15th, 2015. Support for previous versions will be offered for a limited time, but there will no longer be any significant updates to the previous version of H2O.
The following information and links will inform you about what’s new and different and help you prepare to upgrade to H2O 3.0.
Overall, H2O 3.0 is more stable, elegant, and simplified, with additional capabilities not available in previous versions of H2O.
Algorithm Changes
Most of the algorithms available in previous versions of H2O have been improved in terms of speed and accuracy. Currently available model types include Gradient Boosting Machine, Deep Learning, Generalized Linear Model, K-means, Distributed Random Forest, and Naïve Bayes.
There are a few algorithms that are still being refined to provide these same benefits and will be available in a future version of H2O.
Currently, the following algorithms and associated capabilities are still in development:
- Cross-validation
- Grid search
- Principal Component Analysis (PCA)
- Cox Proportional Hazards (Cox PH)
Check back for updates, as these algorithms will be re-introduced in an improved form in a future version of H2O.
Note: The SpeeDRF model has been removed, as it was originally intended as an optimization for small data only. This optimization will be added to the Distributed Random Forest model automatically for small data in a future version of H2O.
Web UI Changes
Our web UI has been completely overhauled with a much more intuitive interface that is similar to IPython Notebook. Each point-and-click action is translated immediately into an individual workflow script that can be saved for later interactive and offline use. As a result, you can now revise and rerun your workflows easily, and can even add comments and rich media.
For more information, refer to our Getting Started with Flow guide, which comprehensively documents how to use Flow. You can also view this brief video, which provides an overview of Flow in action.
API Users
H2O’s new Python API allows Pythonistas to use H2O in their favorite environment. Using the Python command line or an integrated development environment like IPython Notebook H2O users can control clusters and manage massive datasets quickly.
H2O’s REST API is the basis for the web UI (Flow), as well as the R and Python APIs, and is versioned for stability. It is also easier to understand and use, with full metadata available dynamically from the server, allowing for easier integration by developers.
Java Users
Generated Java REST classes ease REST API use by external programs running in a Java Virtual Machine (JVM).
As in previous versions of H2O, users can export trained models as Java objects for easy integration into JVM applications. H2O is currently the only ML tool that provides this capability, making it the data science tool of choice for enterprise developers.
R Users
If you use H2O primarily in R, be aware that as a result of the improvements to the R package for H2O scripts created using previous versions (Nunes 2.8.6.2 or prior) will require minor revisions to work with H2O 3.0.
To assist our R users in upgrading to H2O 3.0 a “shim” tool has been developed. The shim reviews your script, identifies deprecated or revised parameters and arguments, and suggests replacements.
There is also an R Porting Guide that provides a side-by-side comparison of the algorithms in the previous version of H2O with H2O 3.0. It outlines the new, revised, and deprecated parameters for each algorithm, as well as the changes to the output.
Porting R Scripts
This document outlines how to port R scripts written in previous versions of H2O (Nunes 2.8.6.2 or prior, also known as “H2O Classic”) for compatibility with the new H2O 3.0 API. When upgrading from H2O to H2O 3.0, most functions are the same. However, there are some differences that will need to be resolved when porting any scripts that were originally created using H2O to H2O 3.0.
The original R script for H2O is listed first, followed by the updated script for H2O 3.0.
Some of the parameters have been renamed for consistency. For each algorithm, a table that describes the differences is provided.
For additional assistance within R, enter a question mark before the command (for example, ?h2o.glm).
There is also a “shim” available that will review R scripts created with previous versions of H2O, identify deprecated or renamed parameters, and suggest replacements. For more information, refer to the repo here.
Changes from H2O 2.8 to H2O 3.0
h2o.exec
The h2o.exec command is no longer supported. Any workflows using h2o.exec must be revised to remove this command. If the H2O 3.0 workflow contains any parameters or commands from H2O Classic, errors will result and the workflow will fail.
The purpose of h2o.exec was to wrap expressions so that they could be evaluated in a single \Exec2 call. For example,
h2o.exec(fr[,1] + 2/fr[,3])
and
fr[,1] + 2/fr[,3]
produced the same results in H2O. However, the first example makes a single REST call and uses a single temp object, while the second makes several REST calls and uses several temp objects.
Due to the improved architecture in H2O 3.0, the need to use h2o.exec has been eliminated, as the expression can be processed by R as an “unwrapped” typical R expression.
Currently, the only known exception is when factor is used in conjunction with h2o.exec. For example, h2o.exec(fr$myIntCol <- factor(fr$myIntCol)) would become fr$myIntCol <- as.factor(fr$myIntCol)
Note also that an array is not inside a string:
An int array is [1, 2, 3], not “[1, 2, 3]”.
A String array is [“f00”, “b4r”], not “[\”f00\”, \”b4r\”]”
Only string values are enclosed in double quotation marks (").
h2o.performance
To access any exclusively binomial output, use h2o.performance, optionally with the corresponding accessor. The accessor can only use the model metrics object created by h2o.performance. Each accessor is named for its corresponding field (for example, h2o.AUC, h2o.gini, h2o.F1). h2o.performance supports all current algorithms except for K-Means.
If you specify a data frame as a second parameter, H2O will use the specified data frame for scoring. If you do not specify a second parameter, the training metrics for the model metrics object are used.
xval and validation slots
The xval slot has been removed, as nfolds is not currently supported.
The validation slot has been merged with the model slot.
Principal Components Regression (PCR)
Principal Components Regression (PCR) has also been deprecated. To obtain PCR values, create a Principal Components Analysis (PCA) model, then create a GLM model from the scored data from the PCA model.
Table of Contents
GBM
N-fold cross-validation and grid search will be supported in a future version of H2O 3.0.
Renamed GBM Parameters
The following parameters have been renamed, but retain the same functions:
| H2O Classic Parameter Name | H2O 3.0 Parameter Name |
|---|---|
data |
training_frame |
key |
model_id |
n.trees |
ntrees |
interaction.depth |
max_depth |
n.minobsinnode |
min_rows |
shrinkage |
learn_rate |
n.bins |
nbins |
validation |
validation_frame |
balance.classes |
balance_classes |
max.after.balance.size |
max_after_balance_size |
Deprecated GBM Parameters
The following parameters have been removed:
group_split: Bit-set group splitting of categorical variables is now the default.importance: Variable importances are now computed automatically and displayed in the model output.holdout.fraction: The fraction of the training data to hold out for validation is no longer supported.grid.parallelism: Specifying the number of parallel threads to run during a grid search is no longer supported. Grid search will be supported in a future version of H2O 3.0.
New GBM Parameters
The following parameters have been added:
seed: A random number to control sampling and initialization whenbalance_classesis enabled.score_each_iteration: Display error rate information after each tree in the requested set is built.
GBM Algorithm Comparison
| H2O Classic | H2O 3.0 |
|---|---|
h2o.gbm <- function( |
h2o.gbm <- function( |
x, |
x, |
y, |
y, |
data, |
training_frame, |
key = "", |
model_id, |
distribution = 'multinomial', |
distribution = c("bernoulli", "multinomial", "gaussian"), |
n.trees = 10, |
ntrees = 50 |
interaction.depth = 5, |
max_depth = 5, |
n.minobsinnode = 10, |
min_rows = 10, |
shrinkage = 0.1, |
learn_rate = 0.1, |
n.bins = 20, |
nbins = 20, |
validation, |
validation_frame = NULL, |
balance.classes = FALSE |
balance_classes = FALSE, |
max.after.balance.size = 5, |
max_after_balance_size = 1, |
seed, |
|
score_each_iteration) |
|
group_split = TRUE, |
|
importance = FALSE, |
|
nfolds = 0, |
|
holdout.fraction = 0, |
|
class.sampling.factors = NULL, |
|
grid.parallelism = 1) |
Output
The following table provides the component name in H2O, the corresponding component name in H2O 3.0 (if supported), and the model type (binomial, multinomial, or all). Many components are now included in h2o.performance; for more information, refer to (h2o.performance).
| H2O Classic | H2O 3.0 | Model Type |
|---|---|---|
@model$priorDistribution |
all |
|
@model$params |
@allparameters |
all |
@model$err |
@model$scoring_history |
all |
@model$classification |
all |
|
@model$varimp |
@model$variable_importances |
all |
@model$confusion |
@model$training_metrics$cm$table |
binomial and multinomial |
@model$auc |
@model$training_metrics$AUC |
binomial |
@model$gini |
@model$training_metrics$Gini |
binomial |
@model$best_cutoff |
binomial |
|
@model$F1 |
@model$training_metrics$thresholds_and_metric_scores$f1 |
binomial |
@model$F2 |
@model$training_metrics$thresholds_and_metric_scores$f2 |
binomial |
@model$accuracy |
@model$training_metrics$thresholds_and_metric_scores$accuracy |
binomial |
@model$error |
binomial |
|
@model$precision |
@model$training_metrics$thresholds_and_metric_scores$precision |
binomial |
@model$recall |
@model$training_metrics$thresholds_and_metric_scores$recall |
binomial |
@model$mcc |
@model$training_metrics$thresholds_and_metric_scores$absolute_MCC |
binomial |
@model$max_per_class_err |
currently replaced by @model$training_metrics$thresholds_and_metric_scores$min_per_class_correct |
binomial |
GLM
N-fold cross-validation and grid search will be supported in a future version of H2O 3.0.
Renamed GLM Parameters
The following parameters have been renamed, but retain the same functions:
| H2O Classic Parameter Name | H2O 3.0 Parameter Name |
|---|---|
data |
training_frame |
key |
model_id |
nlambda |
nlambdas |
lambda.min.ratio |
lambda_min_ratio |
iter.max |
max_iterations |
epsilon |
beta_epsilon |
Deprecated GLM Parameters
The following parameters have been removed:
return_all_lambda: A logical value indicating whether to return every model built during the lambda search. (may be re-added)higher_accuracy: For improved accuracy, adjust thebeta_epsilonvalue.strong_rules: Discards predictors likely to have 0 coefficients prior to model building. (may be re-added as enabled by default)intercept: Defines factor columns in the model. (may be re-added)non_negative: Specify a non-negative response. (may be re-added)variable_importances: Variable importances are now computed automatically and displayed in the model output. They have been renamed to Normalized Coefficient Magnitudes.disable_line_search: This parameter has been deprecated, as it was mainly used for testing purposes.offset: Specify a column as an offset. (may be re-added)max_predictors: Stops training the algorithm if the number of predictors exceeds the specified value. (may be re-added)
New GLM Parameters
The following parameters have been added:
validation_frame: Specify the validation dataset.solver: Select IRLSM or LBFGS.
GLM Algorithm Comparison
| H2O Classic | H2O 3.0 |
|---|---|
h2o.glm <- function( |
h2o.startGLMJob <- function( |
x, |
x, |
y, |
y, |
data, |
training_frame, |
key = "", |
model_id, |
validation_frame |
|
iter.max = 100, |
max_iterations = 50, |
epsilon = 1e-4 |
beta_epsilon = 0 |
strong_rules = TRUE, |
|
return_all_lambda = FALSE, |
|
intercept = TRUE, |
|
non_negative = FALSE, |
|
solver = c("IRLSM", "L_BFGS"), |
|
standardize = TRUE, |
standardize = TRUE, |
family, |
family = c("gaussian", "binomial", "poisson", "gamma", "tweedie"), |
link, |
link = c("family_default", "identity", "logit", "log", "inverse", "tweedie"), |
tweedie.p = ifelse(family == "tweedie",1.5, NA_real_) |
tweedie_variance_power = NaN, |
tweedie_link_power = NaN, |
|
alpha = 0.5, |
alpha = 0.5, |
prior = NULL |
prior = 0.0, |
lambda = 1e-5, |
lambda = 1e-05, |
lambda_search = FALSE, |
lambda_search = FALSE, |
nlambda = -1, |
nlambdas = -1, |
lambda.min.ratio = -1, |
lambda_min_ratio = 1.0, |
use_all_factor_levels = FALSE |
use_all_factor_levels = FALSE, |
nfolds = 0, |
nfolds = 0, |
beta_constraints = NULL, |
beta_constraint = NULL) |
higher_accuracy = FALSE, |
|
variable_importances = FALSE, |
|
disable_line_search = FALSE, |
|
offset = NULL, |
|
max_predictors = -1) |
Output
The following table provides the component name in H2O, the corresponding component name in H2O 3.0 (if supported), and the model type (binomial, multinomial, or all). Many components are now included in h2o.performance; for more information, refer to (h2o.performance).
| H2O Classic | H2O 3.0 | Model Type |
|---|---|---|
@model$params |
@allparameters |
all |
@model$coefficients |
@model$coefficients |
all |
@model$nomalized_coefficients |
@model$coefficients_table$norm_coefficients |
all |
@model$rank |
@model$rank |
all |
@model$iter |
@model$iter |
all |
@model$lambda |
all |
|
@model$deviance |
@model$residual_deviance |
all |
@model$null.deviance |
@model$null_deviance |
all |
@model$df.residual |
@model$residual_degrees_of_freedom |
all |
@model$df.null |
@model$null_degrees_of_freedom |
all |
@model$aic |
@model$AIC |
all |
@model$train.err |
binomial |
|
@model$prior |
binomial |
|
@model$thresholds |
@model$threshold |
binomial |
@model$best_threshold |
binomial |
|
@model$auc |
@model$AUC |
binomial |
@model$confusion |
binomial |
K-Means
Renamed K-Means Parameters
The following parameters have been renamed, but retain the same functions:
| H2O Classic Parameter Name | H2O 3.0 Parameter Name |
|---|---|
data |
training_frame |
key |
model_id |
centers |
k |
cols |
x |
iter.max |
max_iterations |
normalize |
standardize |
Note In H2O, the normalize parameter was disabled by default. The standardize parameter is enabled by default in H2O 3.0 to provide more accurate results for datasets containing columns with large values.
New K-Means Parameters
The following parameters have been added:
userhas been added as an additional option for theinitparameter. Using this parameter forces the K-Means algorithm to start at the user-specified points.user_points: Specify starting points for the K-Means algorithm.
K-Means Algorithm Comparison
| H2O Classic | H2O 3.0 |
|---|---|
h2o.kmeans <- function( |
h2o.kmeans <- function( |
data, |
training_frame, |
cols = '', |
x, |
centers, |
k, |
key = "", |
model_id, |
iter.max = 10, |
max_iterations = 1000, |
normalize = FALSE, |
standardize = TRUE, |
init = "none", |
init = c("Furthest","Random", "PlusPlus"), |
seed = 0, |
seed) |
Output
The following table provides the component name in H2O and the corresponding component name in H2O 3.0 (if supported).
| H2O Classic | H2O 3.0 |
|---|---|
@model$params |
@allparameters |
@model$centers |
@model$centers |
@model$tot.withinss |
@model$tot_withinss |
@model$size |
@model$size |
@model$iter |
@model$iterations |
@model$_scoring_history |
|
@model$_model_summary |
Deep Learning
- Renamed Deep Learning Parameters
- Deprecated DL Parameters
- New DL Parameters
- DL Algorithm Comparison
- Output
N-fold cross-validation and grid search will be supported in a future version of H2O 3.0.
Note: If the results in the confusion matrix are incorrect, verify that score_training_samples is equal to 0. By default, only the first 10,000 rows are included.
Renamed Deep Learning Parameters
The following parameters have been renamed, but retain the same functions:
| H2O Classic Parameter Name | H2O 3.0 Parameter Name |
|---|---|
data |
training_frame |
key |
model_id |
validation |
validation_frame |
class.sampling.factors |
class_sampling_factors |
nfolds |
n_folds |
override_with_best_model |
overwrite_with_best_model |
Deprecated DL Parameters
The following parameters have been removed:
classification: Classification is now inferred from the data type.holdout_fraction: Fraction of the training data to hold out for validation.
New DL Parameters
The following parameters have been added:
export_weights_and_biases: An additional option allowing users to export the raw weights and biases as H2O frames.
The following options for the loss parameter have been added:
absolute: Provides strong penalties for mispredictionshuber: Can improve results for regression
DL Algorithm Comparison
| H2O Classic | H2O 3.0 |
|---|---|
h2o.deeplearning <- function(x, |
h2o.deeplearning <- function(x, |
y, |
y, |
data, |
training_frame, |
key = "", |
model_id = "", |
override_with_best_model, |
overwrite_with_best_model = true, |
classification = TRUE, |
|
nfolds = 0, |
n_folds = 0 |
validation, |
validation_frame, |
holdout_fraction = 0, |
|
checkpoint = " " |
checkpoint, |
autoencoder, |
autoencoder = false, |
use_all_factor_levels, |
use_all_factor_levels = true |
activation, |
_activation = c("Rectifier", "Tanh", "TanhWithDropout", "RectifierWithDropout", "Maxout", "MaxoutWithDropout"), |
hidden, |
hidden= c(200, 200), |
epochs, |
epochs = 10.0, |
train_samples_per_iteration, |
train_samples_per_iteration = -2, |
seed, |
_seed, |
adaptive_rate, |
adaptive_rate = true, |
rho, |
rho = 0.99, |
epsilon, |
epsilon = 1e-8, |
rate, |
rate = .005, |
rate_annealing, |
rate_annealing = 1e-6, |
rate_decay, |
rate_decay = 1.0, |
momentum_start, |
momentum_start = 0, |
momentum_ramp, |
momentum_ramp = 1e6, |
momentum_stable, |
momentum_stable = 0, |
nesterov_accelerated_gradient, |
nesterov_accelerated_gradient = true, |
input_dropout_ratio, |
input_dropout_ratio = 0.0, |
hidden_dropout_ratios, |
hidden_dropout_ratios, |
l1, |
l1 = 0.0, |
l2, |
l2 = 0.0, |
max_w2, |
max_w2 = Inf, |
initial_weight_distribution, |
initial_weight_distribution = c("UniformAdaptive","Uniform", "Normal"), |
initial_weight_scale, |
initial_weight_scale = 1.0, |
loss, |
loss = "Automatic", "CrossEntropy", "MeanSquare", "Absolute", "Huber"), |
score_interval, |
score_interval = 5, |
score_training_samples, |
score_training_samples = 10000l, |
score_validation_samples, |
score_validation_samples = 0l, |
score_duty_cycle, |
score_duty_cycle = 0.1, |
classification_stop, |
classification_stop = 0 |
regression_stop, |
regression_stop = 1e-6, |
quiet_mode, |
quiet_mode = false, |
max_confusion_matrix_size, |
max_confusion_matrix_size, |
max_hit_ratio_k, |
max_hit_ratio_k, |
balance_classes, |
balance_classes = false, |
class_sampling_factors, |
class_sampling_factors, |
max_after_balance_size, |
max_after_balance_size, |
score_validation_sampling, |
score_validation_sampling, |
diagnostics, |
diagnostics = true, |
variable_importances, |
variable_importances = false, |
fast_mode, |
fast_mode = true, |
ignore_const_cols, |
ignore_const_cols = true, |
force_load_balance, |
force_load_balance = true, |
replicate_training_data, |
replicate_training_data = true, |
single_node_mode, |
single_node_mode = false, |
shuffle_training_data, |
shuffle_training_data = false, |
sparse, |
sparse = false, |
col_major, |
col_major = false, |
max_categorical_features, |
max_categorical_features = Integer.MAX_VALUE, |
reproducible) |
reproducible=FALSE, |
average_activation |
average_activation = 0, |
sparsity_beta = 0 |
|
export_weights_and_biases=FALSE) |
Output
The following table provides the component name in H2O, the corresponding component name in H2O 3.0 (if supported), and the model type (binomial, multinomial, or all). Many components are now included in h2o.performance; for more information, refer to (h2o.performance).
| H2O Classic | H2O 3.0 | Model Type |
|---|---|---|
@model$priorDistribution |
all |
|
@model$params |
@allparameters |
all |
@model$train_class_error |
@model$training_metrics$MSE |
all |
@model$valid_class_error |
@model$validation_metrics$MSE |
all |
@model$varimp |
@model$_variable_importances |
all |
@model$confusion |
@model$training_metrics$cm$table |
binomial and multinomial |
@model$train_auc |
@model$train_AUC |
binomial |
@model$_validation_metrics |
all |
|
@model$_model_summary |
all |
|
@model$_scoring_history |
all |
Distributed Random Forest
- Changes to DRF in H2O 3.0
- Renamed DRF Parameters
- Deprecated DRF Parameters
- New DRF Parameters
- DRF Algorithm Comparison
- Output
Changes to DRF in H2O 3.0
Distributed Random Forest (DRF) was represented as h2o.randomForest(type="BigData", ...) in H2O Classic. In H2O Classic, SpeeDRF (type="fast") was not as accurate, especially for complex data with categoricals, and did not address regression problems. DRF (type="BigData") was at least as accurate as SpeeDRT (type="fast") and was the only algorithm that scaled to big data (data too large to fit on a single node).
In H2O 3.0, our plan is to improve the performance of DRF so that the data fits on a single node (optimally, for all cases), which will make SpeeDRF obsolete. Ultimately, the goal is provide a single algorithm that provides the “best of both worlds” for all datasets and use cases.
Note: H2O 3.0 only supports DRF. SpeeDRF is no longer supported. The functionality of DRF in H2O 3.0 is similar to DRF functionality in H2O.
Renamed DRF Parameters
The following parameters have been renamed, but retain the same functions:
| H2O Classic Parameter Name | H2O 3.0 Parameter Name |
|---|---|
data |
training_frame |
key |
model_id |
validation |
validation_frame |
sample.rate |
sample_rate |
ntree |
ntrees |
depth |
max_depth |
balance.classes |
balance_classes |
score.each.iteration |
score_each_iteration |
class.sampling.factors |
class_sampling_factors |
nodesize |
min_rows |
Deprecated DRF Parameters
The following parameters have been removed:
classification: This is now automatically inferred from the response type. To achieve classification with a 0/1 response column, explicitly convert the response to a factor (as.factor()).importance: Variable importances are now computed automatically and displayed in the model output.holdout.fraction: Specifying the fraction of the training data to hold out for validation is no longer supported.doGrpSplit: The bit-set group splitting of categorical variables is now the default.verbose: Infonrmation about tree splits and extra statistics is now included automatically in the stdout.oobee: The out-of-bag error estimate is now computed automatically (if no validation set is specified).stat.type: This parameter was used for SpeeDRF, which is no longer supported.type: This parameter was used for SpeeDRF, which is no longer supported.
New DRF Parameters
The following parameter has been added:
build_tree_one_node: Run on a single node to use fewer CPUs.
DRF Algorithm Comparison
| H2O Classic | H2O 3.0 |
|---|---|
h2o.randomForest <- function(x, |
h2o.randomForest <- function( |
x, |
x, |
y, |
y, |
data, |
training_frame, |
key="", |
model_id, |
validation, |
validation_frame, |
mtries = -1, |
mtries = -1, |
sample.rate=2/3, |
sample_rate = 0.632, |
build_tree_one_node = FALSE, |
|
ntree=50 |
ntrees=50, |
depth=20, |
max_depth = 20, |
min_rows = 1, |
|
nbins=20, |
nbins = 20, |
balance.classes = FALSE, |
balance_classes = FALSE, |
score.each.iteration = FALSE, |
score_each_iteration = FALSE, |
seed = -1, |
seed |
nodesize = 1, |
|
classification=TRUE, |
|
importance=FALSE, |
|
nfolds=0, |
|
holdout.fraction = 0, |
|
max.after.balance.size = 5, |
max_after_balance_size) |
class.sampling.factors = NULL, |
|
doGrpSplit = TRUE, |
|
verbose = FALSE, |
|
oobee = TRUE, |
|
stat.type = "ENTROPY", |
|
type = "fast") |
Output
The following table provides the component name in H2O, the corresponding component name in H2O 3.0 (if supported), and the model type (binomial, multinomial, or all). Many components are now included in h2o.performance; for more information, refer to (h2o.performance).
| H2O Classic | H2O 3.0 | Model Type |
|---|---|---|
@model$priorDistribution |
all |
|
@model$params |
@allparameters |
all |
@model$mse |
@model$scoring_history |
all |
@model$forest |
@model$model_summary |
all |
@model$classification |
all |
|
@model$varimp |
@model$variable_importances |
all |
@model$confusion |
@model$training_metrics$cm$table |
binomial and multinomial |
@model$auc |
@model$training_metrics$AUC |
binomial |
@model$gini |
@model$training_metrics$Gini |
binomial |
@model$best_cutoff |
binomial |
|
@model$F1 |
@model$training_metrics$thresholds_and_metric_scores$f1 |
binomial |
@model$F2 |
@model$training_metrics$thresholds_and_metric_scores$f2 |
binomial |
@model$accuracy |
@model$training_metrics$thresholds_and_metric_scores$accuracy |
binomial |
@model$Error |
@model$Error |
binomial |
@model$precision |
@model$training_metrics$thresholds_and_metric_scores$precision |
binomial |
@model$recall |
@model$training_metrics$thresholds_and_metric_scores$recall |
binomial |
@model$mcc |
@model$training_metrics$thresholds_and_metric_scores$absolute_MCC |
binomial |
@model$max_per_class_err |
currently replaced by @model$training_metrics$thresholds_and_metric_scores$min_per_class_correct |
binomial |
Github Users
All users who pull directly from the H2O classic repo on Github should be aware that this repo will be renamed. To retain access to the original H2O (2.8.6.2 and prior) repository:
The simple way
This is the easiest way to change your local repo and is recommended for most users.
- Enter
git remote -vto view a list of your repositories. Copy the address your H2O classic repo (refer to the text in brackets below - your address will vary depending on your connection method):
H2O_User-MBP:h2o H2O_User$ git remote -v origin https://{H2O_User@github.com}/h2oai/h2o.git (fetch) origin https://{H2O_User@github.com}/h2oai/h2o.git (push)- Enter
git remote set-url origin {H2O_User@github.com}:h2oai/h2o-2.git, where{H2O_User@github.com}represents the address copied in the previous step.
The more complicated way
This method involves editing the Github config file and should only be attempted by users who are confident enough with their knowledge of Github to do so.
- Enter
vim .git/config. Look for the
[remote "origin"]section:[remote "origin"] url = https://H2O_User@github.com/h2oai/h2o.git fetch = +refs/heads/*:refs/remotes/origin/*- In the
url =line, changeh2o.gittoh2o-2.git. - Save the changes.
The latest version of H2O is stored in the h2o-3 repository. All previous links to this repo will still work, but if you would like to manually update your Github configuration, follow the instructions above, replacing h2o-2 with h2o-3.
FAQ
How do I score using an exported JSON model?
Since JSON is just a representation format, it cannot be directly executed, so a JSON export can’t be used for scoring. However, you can score by:
- including the POJO in your execution stream and handing it observations one at a time
or
- handing your data in bulk to an H2O cluster, which will score using high throughput parallel and distributed bulk scoring.
How do I predict using multiple response variables?
Currently, H2O does not support multiple response variables. To predict different response variables, build multiple modes.
How do I kill any running instances of H2O?
In Terminal, enter ps -efww | grep h2o, then kill any running PIDs.
Why is H2O not launching from the command line?
$ java -jar h2o.jar &
% Exception in thread "main" java.lang.ExceptionInInitializerError
at java.lang.Class.initializeClass(libgcj.so.10)
at water.Boot.getMD5(Boot.java:73)
at water.Boot.<init>(Boot.java:114)
at water.Boot.<clinit>(Boot.java:57)
at java.lang.Class.initializeClass(libgcj.so.10)
Caused by: java.lang.IllegalArgumentException
at java.util.regex.Pattern.compile(libgcj.so.10)
at water.util.Utils.<clinit>(Utils.java:1286)
at java.lang.Class.initializeClass(libgcj.so.10)
...4 more
The only prerequisite for running H2O is a compatible version of Java. We recommend Oracle's Java 1.7 <http://www.oracle.com/technetwork/java/javase/downloads/jdk7-downloads-1880260.html>_.
Why did I receive the following error when I tried to launch H2O?
[root@sandbox h2o-dev-0.3.0.1188-hdp2.2]hadoop jar h2odriver.jar -nodes 2 -mapperXmx 1g -output hdfsOutputDirName
Determining driver host interface for mapper->driver callback...
[Possible callback IP address: 10.0.2.15]
[Possible callback IP address: 127.0.0.1]
Using mapper->driver callback IP address and port: 10.0.2.15:41188
(You can override these with -driverif and -driverport.)
Memory Settings:
mapreduce.map.java.opts: -Xms1g -Xmx1g -Dlog4j.defaultInitOverride=true
Extra memory percent: 10
mapreduce.map.memory.mb: 1126
15/05/08 02:33:40 INFO impl.TimelineClientImpl: Timeline service address: http://sandbox.hortonworks.com:8188/ws/v1/timeline/
15/05/08 02:33:41 INFO client.RMProxy: Connecting to ResourceManager at sandbox.hortonworks.com/10.0.2.15:8050
15/05/08 02:33:47 INFO mapreduce.JobSubmitter: number of splits:2
15/05/08 02:33:48 INFO mapreduce.JobSubmitter: Submitting tokens for job: job_1431052132967_0001
15/05/08 02:33:51 INFO impl.YarnClientImpl: Submitted application application_1431052132967_0001
15/05/08 02:33:51 INFO mapreduce.Job: The url to track the job: http://sandbox.hortonworks.com:8088/proxy/application_1431052132967_0001/
Job name 'H2O_3889' submitted
JobTracker job ID is 'job_1431052132967_0001'
For YARN users, logs command is 'yarn logs -applicationId application_1431052132967_0001'
Waiting for H2O cluster to come up...
H2O node 10.0.2.15:54321 requested flatfile
ERROR: Timed out waiting for H2O cluster to come up (120 seconds)
ERROR: (Try specifying the -timeout option to increase the waiting time limit)
15/05/08 02:35:59 INFO impl.TimelineClientImpl: Timeline service address: http://sandbox.hortonworks.com:8188/ws/v1/timeline/
15/05/08 02:35:59 INFO client.RMProxy: Connecting to ResourceManager at sandbox.hortonworks.com/10.0.2.15:8050
----- YARN cluster metrics -----
Number of YARN worker nodes: 1
----- Nodes -----
Node: http://sandbox.hortonworks.com:8042 Rack: /default-rack, RUNNING, 1 containers used, 0.2 / 2.2 GB used, 1 / 8 vcores used
----- Queues -----
Queue name: default
Queue state: RUNNING
Current capacity: 0.11
Capacity: 1.00
Maximum capacity: 1.00
Application count: 1
----- Applications in this queue -----
Application ID: application_1431052132967_0001 (H2O_3889)
Started: root (Fri May 08 02:33:50 UTC 2015)
Application state: FINISHED
Tracking URL: http://sandbox.hortonworks.com:8088/proxy/application_1431052132967_0001/jobhistory/job/job_1431052132967_0001
Queue name: default
Used/Reserved containers: 1 / 0
Needed/Used/Reserved memory: 0.2 GB / 0.2 GB / 0.0 GB
Needed/Used/Reserved vcores: 1 / 1 / 0
Queue 'default' approximate utilization: 0.2 / 2.2 GB used, 1 / 8 vcores used
----------------------------------------------------------------------
ERROR: Job memory request (2.2 GB) exceeds available YARN cluster memory (2.2 GB)
WARNING: Job memory request (2.2 GB) exceeds queue available memory capacity (2.0 GB)
ERROR: Only 1 out of the requested 2 worker containers were started due to YARN cluster resource limitations
----------------------------------------------------------------------
Attempting to clean up hadoop job...
15/05/08 02:35:59 INFO impl.YarnClientImpl: Killed application application_1431052132967_0001
Killed.
[root@sandbox h2o-dev-0.3.0.1188-hdp2.2]#
The H2O launch failed because more memory was requested than was available. Make sure you are not trying to specify more memory in the launch parameters than you have available.
Clusters
When trying to launch H2O, I received the following error message: ERROR: Too many retries starting cloud. What should I do?
If you are trying to start a multi-node cluster where the nodes use multiple network interfaces, by default H2O will resort to using the default host (127.0.0.1).
To specify an IP address, launch H2O using the following command:
java -jar h2o.jar -ip <IP_Address> -port <PortNumber>
If this does not resolve the issue, try the following additional troubleshooting tips:
- Test connectivity using
curl: First, log in to the first node and entercurl http://<Node2IP>:54321(where<Node2IP>is the IP address of the second node. Then, log in to the second node and entercurl http://<Node1IP>:54321(where<Node1IP>is the IP address of the first node). Look for output from H2O. - Confirm ports 54321 and 54322 are available for both TCP and UDP.
- Confirm your firewall is not preventing the nodes from locating each other.
- Check if you have SELINUX or IPTABLES enabled; if so, disable them.
- Check the configuration for the EC2 security group.
- Confirm that the username is the same on all nodes; if not, define the cloud using
-name. - Check if the nodes are on different networks.
- Check if the nodes have different interfaces; if so, use the
-networkoption to define the network (for example,-network 127.0.0.1). - Force the bind address using
-ip. - Confirm the nodes are not using different versions of H2O.
What should I do if I tried to start a cluster but the nodes started independent clouds that are not connected?
Because the default cloud name is the user name of the node, if the nodes are on different operating systems (for example, one node is using Windows and the other uses OS X), the different user names on each machine will prevent the nodes from recognizing that they belong to the same cloud. To resolve this issue, use -name to configure the same name for all nodes.
One of the nodes in my cluster is unavailable - what do I do?
H2O does not support high availability (HA). If a node in the cluster is unavailable, bring the cluster down and create a new healthy cluster.
How do I add new nodes to an existing cluster?
New nodes can only be added if H2O has not started any jobs. Once H2O starts a task, it locks the cluster to prevent new nodes from joining. If H2O has started a job, you must create a new cluster to include additional nodes.
How do I check if all the nodes in the cluster are healthy and communicating?
In the Flow web UI, click the Admin menu and select Cluster Status.
How do I create a cluster behind a firewall?
H2O uses two ports:
- The
REST_APIport (54321): Specify when launching H2O using-port; uses TCP only. - The
INTERNAL_COMMUNICATIONport (54322): Implied based on the port specified as theREST_APIport, +1; requires TCP and UDP.
You can start the cluster behind the firewall, but to reach it, you must make a tunnel to reach the REST_API port. To use the cluster, the REST_API port of at least one node must be reachable.
How do I specify which nodes should run H2O in a Hadoop cluster?
Currently, this is not yet supported. To provide resource isolation (for example, to isolate H2O to the worker nodes, rather than the master nodes), use YARN Nodemanagers to specify the nodes to use.
I launched H2O instances on my nodes - why won’t they form a cloud?
If you launch without specifying the IP address by adding argument -ip:
$ java -Xmx20g -jar h2o.jar -flatfile flatfile.txt -port 54321
and multiple local IP addresses are detected, H2O uses the default localhost (127.0.0.1) as shown below:
10:26:32.266 main WARN WATER: Multiple local IPs detected:
+ /198.168.1.161 /198.168.58.102
+ Attempting to determine correct address...
10:26:32.284 main WARN WATER: Failed to determine IP, falling back to localhost.
10:26:32.325 main INFO WATER: Internal communication uses port: 54322
+ Listening for HTTP and REST traffic
+ on http://127.0.0.1:54321/
10:26:32.378 main WARN WATER: Flatfile configuration does not include self:
/127.0.0.1:54321 but contains [/192.168.1.161:54321, /192.168.1.162:54321]
To avoid using 127.0.0.1 on servers with multiple local IP addresses, run the command with the -ip argument to force H2O to launch at the specified IP:
$ java -Xmx20g -jar h2o.jar -flatfile flatfile.txt -ip 192.168.1.161 -port 54321
Sparkling Water
How do I inspect H2O using Flow while a droplet is running?
If your droplet execution time is very short, add a simple sleep statement to your code:
Thread.sleep(...)
How do I change the memory size of the executors in a droplet?
There are two ways to do this:
Change your default Spark setup in
$SPARK_HOME/conf/spark-defaults.confor
Pass
--confvia spark-submit when you launch your droplet (e.g.,$SPARK_HOME/bin/spark-submit --conf spark.executor.memory=4g --master $MASTER --class org.my.Droplet $TOPDIR/assembly/build/libs/droplet.jar
Tunneling between servers with H2O
To tunnel between servers (for example, due to firewalls):
- Use ssh to log in to the machine where H2O will run.
Start an instance of H2O by locating the working directory and calling a java command similar to the following example.
The port number chosen here is arbitrary; yours may be different.
$ java -jar h2o.jar -port 55599This returns output similar to the following:
irene@mr-0x3:~/target$ java -jar h2o.jar -port 55599 04:48:58.053 main INFO WATER: ----- H2O started ----- 04:48:58.055 main INFO WATER: Build git branch: master 04:48:58.055 main INFO WATER: Build git hash: 64fe68c59ced5875ac6bac26a784ce210ef9f7a0 04:48:58.055 main INFO WATER: Build git describe: 64fe68c 04:48:58.055 main INFO WATER: Build project version: 1.7.0.99999 04:48:58.055 main INFO WATER: Built by: 'Irene' 04:48:58.055 main INFO WATER: Built on: 'Wed Sep 4 07:30:45 PDT 2013' 04:48:58.055 main INFO WATER: Java availableProcessors: 4 04:48:58.059 main INFO WATER: Java heap totalMemory: 0.47 gb 04:48:58.059 main INFO WATER: Java heap maxMemory: 6.96 gb 04:48:58.060 main INFO WATER: ICE root: '/tmp' 04:48:58.081 main INFO WATER: Internal communication uses port: 55600 + Listening for HTTP and REST traffic on + http://192.168.1.173:55599/ 04:48:58.109 main INFO WATER: H2O cloud name: 'irene' 04:48:58.109 main INFO WATER: (v1.7.0.99999) 'irene' on /192.168.1.173:55599, discovery address /230 .252.255.19:59132 04:48:58.111 main INFO WATER: Cloud of size 1 formed [/192.168.1.173:55599] 04:48:58.247 main INFO WATER: Log dir: '/tmp/h2ologs'Log into the remote machine where the running instance of H2O will be forwarded using a command similar to the following (your specified port numbers and IP address will be different)
ssh -L 55577:localhost:55599 irene@192.168.1.173Check the cluster status.
You are now using H2O from localhost:55577, but the instance of H2O is running on the remote server (in this case the server with the ip address 192.168.1.xxx) at port number 55599.
To see this in action note that the web UI is pointed at localhost:55577, but that the cluster status shows the cluster running on 192.168.1.173:55599
Quick Start Videos
- H2O Quick Start with Flow
- H2O Quick Start with Python
- H2O Quick Start on Hadoop
- H2O Quick Start with Sparkling Water
- H2O Quick Start with R
H2O Quick Start with Flow
H2O Quick Start with Python
H2O Quick Start on Hadoop
H2O Quick Start with Sparkling Water
H2O Quick Start with R
REST API Reference
- /3/About
- /3/Cloud
- /3/Cloud
- /3/Configuration/ModelBuilders/visibility
- /3/Configuration/ModelBuilders/visibility
- /3/CreateFrame
- /3/DKV
- /3/DKV/(?
.*) - /3/DownloadDataset
- /3/Find
- /3/Frames
- /3/Frames
- /3/Frames/(?
.*) - /3/Frames/(?
.*) - /3/Frames/(?
.*)/columns - /3/Frames/(?
.*)/columns/(? .*) - /3/Frames/(?
.*)/columns/(? .*)/domain - /3/Frames/(?
.*)/columns/(? .*)/summary - /3/Frames/(?
.*)/export/(? .*)/overwrite/(? .*) - /3/Frames/(?
.*)/summary - /3/GarbageCollect
- /3/Grid/drf
- /3/Grid/gbm
- /3/Grid/kmeans
- /3/ImportFiles
- /3/InitID
- /3/Interaction
- /3/JStack
- /3/Jobs
- /3/Jobs/(?
.*) - /3/Jobs/(?
.*)/cancel - /3/KillMinus3
- /3/LogAndEcho
- /3/Logs/nodes/(?
.*)/files/(? .*) - /3/MakeGLMModel
- /3/Metadata/endpoints
- /3/Metadata/endpoints/(?
[0-9]+) - /3/Metadata/endpoints/(?
.*) - /3/Metadata/schemaclasses/(?
.*) - /3/Metadata/schemas
- /3/Metadata/schemas/(?
.*) - /3/MissingInserter
- /3/ModelBuilders
- /3/ModelBuilders/(?
.*) - /3/ModelBuilders/deeplearning
- /3/ModelBuilders/deeplearning/parameters
- /3/ModelBuilders/drf
- /3/ModelBuilders/drf/parameters
- /3/ModelBuilders/gbm
- /3/ModelBuilders/gbm/parameters
- /3/ModelBuilders/glm
- /3/ModelBuilders/glm/parameters
- /3/ModelBuilders/glrm
- /3/ModelBuilders/glrm/parameters
- /3/ModelBuilders/kmeans
- /3/ModelBuilders/kmeans/parameters
- /3/ModelBuilders/naivebayes
- /3/ModelBuilders/naivebayes/parameters
- /3/ModelBuilders/pca
- /3/ModelBuilders/pca/parameters
- /3/ModelBuilders/svd
- /3/ModelBuilders/svd/parameters
- /3/ModelMetrics
- /3/ModelMetrics/frames/(?.*)
- /3/ModelMetrics/frames/(?.*)/models/(?
.*) - /3/ModelMetrics/frames/(?.*)/models/(?
.*) - /3/ModelMetrics/models/(?
.*) - /3/ModelMetrics/models/(?
.*)/frames/(?.*) - /3/ModelMetrics/models/(?
.*)/frames/(?.*) - /3/ModelMetrics/models/(?
.*)/frames/(?.*) - /3/Models
- /3/Models
- /3/Models/(?
.*) - /3/Models/(?
.*) - /3/Models/(?
.*)/preview - /3/NetworkTest
- /3/NodePersistentStorage/(?
.*) - /3/NodePersistentStorage/(?
.*) - /3/NodePersistentStorage/(?
.*)/(? .*) - /3/NodePersistentStorage/(?
.*)/(? .*) - /3/NodePersistentStorage/(?
.*)/(? .*) - /3/NodePersistentStorage/categories/(?
.*)/exists - /3/NodePersistentStorage/categories/(?
.*)/names/(? .*)/exists - /3/NodePersistentStorage/configured
- /3/Parse
- /3/ParseSetup
- /3/Predictions/models/(?
.*)/frames/(?.*) - /3/Profiler
- /3/Rapids
- /3/Rapids/isEval
- /3/Shutdown
- /3/SplitFrame
- /3/Timeline
- /3/Tutorials
- /3/Typeahead/files
- /3/UnlockKeys
- /3/WaterMeterCpuTicks/(?
.*) - /3/WaterMeterIo
- /3/WaterMeterIo/(?
.*) - /99/Sample
GET /3/About
Return information about this H2O.
| Input | AboutV3 |
| Output | AboutV3 |
GET /3/Cloud
Determine the status of the nodes in the H2O cloud.
| Input | CloudV3 |
| Output | CloudV3 |
HEAD /3/Cloud
Determine the status of the nodes in the H2O cloud.
| Input | CloudV3 |
| Output | CloudV3 |
POST /3/Configuration/ModelBuilders/visibility
Set Model Builders visibility level.
| Input | ModelBuildersVisibilityV3 |
| Output | ModelBuildersVisibilityV3 |
GET /3/Configuration/ModelBuilders/visibility
Get Model Builders visibility level.
| Input | ModelBuildersVisibilityV3 |
| Output | ModelBuildersVisibilityV3 |
POST /3/CreateFrame
Create a synthetic H2O Frame.
| Input | CreateFrameV3 |
| Output | CreateFrameV3 |
DELETE /3/DKV
Remove all keys from the H2O distributed K/V store.
| Input | RemoveAllV3 |
| Output | RemoveAllV3 |
DELETE /3/DKV/(?.*)
Remove an arbitrary key from the H2O distributed K/V store.
| Input | RemoveV3 |
| Output | RemoveV3 |
GET /3/DownloadDataset
Download something something.
| Input | DownloadDataV3 |
| Output | DownloadDataV3 |
GET /3/Find
Find a value within a Frame.
| Input | FindV3 |
| Output | FindV3 |
GET /3/Frames
Return all Frames in the H2O distributed K/V store.
| Input | FramesV3 |
| Output | FramesV3 |
DELETE /3/Frames
Delete all Frames from the H2O distributed K/V store.
| Input | FramesV3 |
| Output | FramesV3 |
GET /3/Frames/(?.*)
Return the specified Frame.
| Input | FramesV3 |
| Output | FramesV3 |
DELETE /3/Frames/(?.*)
Delete the specified Frame from the H2O distributed K/V store.
| Input | FramesV3 |
| Output | FramesV3 |
GET /3/Frames/(?.*)/columns
Return all the columns from a Frame.
| Input | FramesV3 |
| Output | FramesV3 |
GET /3/Frames/(?.*)/columns/(?.*)
Return the specified column from a Frame.
| Input | FramesV3 |
| Output | FramesV3 |
GET /3/Frames/(?.*)/columns/(?.*)/domain
Return the domains for the specified column. “null” if the column is not an Enum.
| Input | FramesV3 |
| Output | FramesV3 |
GET /3/Frames/(?.*)/columns/(?.*)/summary
Return the summary metrics for a column, e.g. mins, maxes, mean, sigma, percentiles, etc.
| Input | FramesV3 |
| Output | FramesV3 |
GET /3/Frames/(?.*)/export/(?.*)/overwrite/(?.*)
Export a Frame to the given path with optional overwrite.
| Input | FramesV3 |
| Output | FramesV3 |
GET /3/Frames/(?.*)/summary
Return a Frame, including the histograms, after forcing computation of rollups.
| Input | FramesV3 |
| Output | FramesV3 |
POST /3/GarbageCollect
Explicitly call System.gc().
| Input | GarbageCollectV3 |
| Output | GarbageCollectV3 |
POST /3/Grid/drf
Run grid search for DRF model.
| Input | DRFGridSearchV3 |
| Output | DRFGridSearchV3 |
POST /3/Grid/gbm
Run grid search for GBM model.
| Input | GBMGridSearchV3 |
| Output | GBMGridSearchV3 |
POST /3/Grid/kmeans
Run grid search for KMeans model.
| Input | KMeansGridSearchV3 |
| Output | KMeansGridSearchV3 |
GET /3/ImportFiles
Import raw data files into a single-column H2O Frame.
| Input | ImportFilesV3 |
| Output | ImportFilesV3 |
GET /3/InitID
Issue a new session ID.
| Input | InitIDV3 |
| Output | InitIDV3 |
POST /3/Interaction
Create interactions between categorical columns.
| Input | InteractionV3 |
| Output | InteractionV3 |
GET /3/JStack
Something something something.
| Input | JStackV3 |
| Output | JStackV3 |
GET /3/Jobs
Get a list of all the H2O Jobs (long-running actions).
| Input | JobsV3 |
| Output | Schema |
GET /3/Jobs/(?.*)
Get the status of the given H2O Job (long-running action).
| Input | JobsV3 |
| Output | Schema |
POST /3/Jobs/(?.*)/cancel
Cancel a running job.
| Input | JobsV3 |
| Output | Schema |
GET /3/KillMinus3
Kill minus 3 on this node
| Input | KillMinus3V3 |
| Output | KillMinus3V3 |
POST /3/LogAndEcho
Save a message to the H2O logfile.
| Input | LogAndEchoV3 |
| Output | LogAndEchoV3 |
GET /3/Logs/nodes/(?.*)/files/(?.*)
Get named log file for a node.
| Input | LogsV3 |
| Output | LogsV3 |
POST /3/MakeGLMModel
make a new GLM model based on existing one
| Input | MakeGLMModelV3 |
| Output | GLMModelV3 |
GET /3/Metadata/endpoints
Return a list of all the REST API endpoints.
| Input | MetadataV3 |
| Output | MetadataV3 |
GET /3/Metadata/endpoints/(?[0-9]+)
Return the REST API endpoint metadata, including documentation, for the endpoint specified by number.
| Input | MetadataV3 |
| Output | MetadataV3 |
GET /3/Metadata/endpoints/(?.*)
Return the REST API endpoint metadata, including documentation, for the endpoint specified by path.
| Input | MetadataV3 |
| Output | MetadataV3 |
GET /3/Metadata/schemaclasses/(?.*)
Return the REST API schema metadata for specified schema class.
| Input | MetadataV3 |
| Output | MetadataV3 |
GET /3/Metadata/schemas
Return list of all REST API schemas.
| Input | MetadataV3 |
| Output | MetadataV3 |
GET /3/Metadata/schemas/(?.*)
Return the REST API schema metadata for specified schema.
| Input | MetadataV3 |
| Output | MetadataV3 |
POST /3/MissingInserter
Insert missing values.
| Input | MissingInserterV3 |
| Output | MissingInserterV3 |
GET /3/ModelBuilders
Return the Model Builder metadata for all available algorithms.
| Input | ModelBuildersV3 |
| Output | ModelBuildersV3 |
GET /3/ModelBuilders/(?.*)
Return the Model Builder metadata for the specified algorithm.
| Input | ModelBuildersV3 |
| Output | ModelBuildersV3 |
POST /3/ModelBuilders/deeplearning
Train a Deep Learning model on the specified Frame.
| Input | DeepLearningV3 |
| Output | Schema |
POST /3/ModelBuilders/deeplearning/parameters
Validate a set of Deep Learning model builder parameters.
| Input | DeepLearningV3 |
| Output | DeepLearningV3 |
POST /3/ModelBuilders/drf
Train a DRF model on the specified Frame.
| Input | DRFV3 |
| Output | Schema |
POST /3/ModelBuilders/drf/parameters
Validate a set of DRF model builder parameters.
| Input | DRFV3 |
| Output | DRFV3 |
POST /3/ModelBuilders/gbm
Train a GBM model on the specified Frame.
| Input | GBMV3 |
| Output | Schema |
POST /3/ModelBuilders/gbm/parameters
Validate a set of GBM model builder parameters.
| Input | GBMV3 |
| Output | GBMV3 |
POST /3/ModelBuilders/glm
Train a GLM model on the specified Frame.
| Input | GLMV3 |
| Output | Schema |
POST /3/ModelBuilders/glm/parameters
Validate a set of GLM model builder parameters.
| Input | GLMV3 |
| Output | GLMV3 |
POST /3/ModelBuilders/glrm
Train a GLRM model on the specified Frame.
| Input | GLRMV3 |
| Output | Schema |
POST /3/ModelBuilders/glrm/parameters
Validate a set of GLRM model builder parameters.
| Input | GLRMV3 |
| Output | GLRMV3 |
POST /3/ModelBuilders/kmeans
Train a KMeans model on the specified Frame.
| Input | KMeansV3 |
| Output | Schema |
POST /3/ModelBuilders/kmeans/parameters
Validate a set of KMeans model builder parameters.
| Input | KMeansV3 |
| Output | KMeansV3 |
POST /3/ModelBuilders/naivebayes
Train a Naive Bayes model on the specified Frame.
| Input | NaiveBayesV3 |
| Output | Schema |
POST /3/ModelBuilders/naivebayes/parameters
Validate a set of Naive Bayes model builder parameters.
| Input | NaiveBayesV3 |
| Output | NaiveBayesV3 |
POST /3/ModelBuilders/pca
Train a PCA model on the specified Frame.
| Input | PCAV3 |
| Output | Schema |
POST /3/ModelBuilders/pca/parameters
Validate a set of PCA model builder parameters.
| Input | PCAV3 |
| Output | PCAV3 |
POST /3/ModelBuilders/svd
Train a SVD model on the specified Frame.
| Input | SVDV3 |
| Output | Schema |
POST /3/ModelBuilders/svd/parameters
Validate a set of SVD model builder parameters.
| Input | SVDV3 |
| Output | SVDV3 |
GET /3/ModelMetrics
Return all the saved scoring metrics.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
GET /3/ModelMetrics/frames/(?.*)
Return the saved scoring metrics for the specified Frame.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
GET /3/ModelMetrics/frames/(?.*)/models/(?.*)
Return the saved scoring metrics for the specified Model and Frame.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
DELETE /3/ModelMetrics/frames/(?.*)/models/(?.*)
Return the saved scoring metrics for the specified Model and Frame.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
GET /3/ModelMetrics/models/(?.*)
Return the saved scoring metrics for the specified Model.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
GET /3/ModelMetrics/models/(?.*)/frames/(?.*)
Return the saved scoring metrics for the specified Model and Frame.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
DELETE /3/ModelMetrics/models/(?.*)/frames/(?.*)
Return the saved scoring metrics for the specified Model and Frame.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
POST /3/ModelMetrics/models/(?.*)/frames/(?.*)
Return the scoring metrics for the specified Frame with the specified Model. If the Frame has already been scored with the Model then cached results will be returned; otherwise predictions for all rows in the Frame will be generated and the metrics will be returned.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
GET /3/Models
Return all Models from the H2O distributed K/V store.
| Input | ModelsV3 |
| Output | ModelsV3 |
DELETE /3/Models
Delete all Models from the H2O distributed K/V store.
| Input | ModelsV3 |
| Output | ModelsV3 |
GET /3/Models/(?.*)
Return the specified Model from the H2O distributed K/V store, optionally with the list of compatible Frames.
| Input | ModelsV3 |
| Output | ModelsV3 |
DELETE /3/Models/(?.*)
Delete the specified Model from the H2O distributed K/V store.
| Input | ModelsV3 |
| Output | ModelsV3 |
GET /3/Models/(?.*)/preview
Return potentially abridged model suitable for viewing in a browser (currently only used for java model code).
| Input | ModelsV3 |
| Output | ModelsV3 |
GET /3/NetworkTest
Something something something.
| Input | NetworkTestV3 |
| Output | NetworkTestV3 |
POST /3/NodePersistentStorage/(?.*)
Store a value.
| Input | NodePersistentStorageV3 |
| Output | NodePersistentStorageV3 |
GET /3/NodePersistentStorage/(?.*)
Return all keys stored for a given category.
| Input | NodePersistentStorageV3 |
| Output | NodePersistentStorageV3 |
POST /3/NodePersistentStorage/(?.*)/(?.*)
Store a named value.
| Input | NodePersistentStorageV3 |
| Output | NodePersistentStorageV3 |
GET /3/NodePersistentStorage/(?.*)/(?.*)
Return value for a given name.
| Input | NodePersistentStorageV3 |
| Output | NodePersistentStorageV3 |
DELETE /3/NodePersistentStorage/(?.*)/(?.*)
Delete a key.
| Input | NodePersistentStorageV3 |
| Output | NodePersistentStorageV3 |
GET /3/NodePersistentStorage/categories/(?.*)/exists
Return true or false.
| Input | NodePersistentStorageV3 |
| Output | NodePersistentStorageV3 |
GET /3/NodePersistentStorage/categories/(?.*)/names/(?.*)/exists
Return true or false.
| Input | NodePersistentStorageV3 |
| Output | NodePersistentStorageV3 |
GET /3/NodePersistentStorage/configured
Return true or false.
| Input | NodePersistentStorageV3 |
| Output | NodePersistentStorageV3 |
POST /3/Parse
Parse a raw byte-oriented Frame into a useful columnar data Frame.
| Input | ParseV3 |
| Output | ParseV3 |
POST /3/ParseSetup
Guess the parameters for parsing raw byte-oriented data into an H2O Frame.
| Input | ParseSetupV3 |
| Output | ParseSetupV3 |
POST /3/Predictions/models/(?.*)/frames/(?.*)
Score (generate predictions) for the specified Frame with the specified Model. Both the Frame of predictions and the metrics will be returned.
| Input | ModelMetricsListSchemaV3 |
| Output | ModelMetricsListSchemaV3 |
GET /3/Profiler
Something something something.
| Input | ProfilerV3 |
| Output | ProfilerV3 |
POST /3/Rapids
Something something R exec something.
| Input | RapidsV3 |
| Output | RapidsV3 |
GET /3/Rapids/isEval
something something r exec something.
| Input | RapidsV3 |
| Output | RapidsV3 |
POST /3/Shutdown
Shut down the cluster
| Input | ShutdownV3 |
| Output | ShutdownV3 |
POST /3/SplitFrame
Split a H2O Frame.
| Input | SplitFrameV3 |
| Output | SplitFrameV3 |
GET /3/Timeline
Something something something.
| Input | TimelineV3 |
| Output | TimelineV3 |
GET /3/Tutorials
H2O tutorials.
| Input | TutorialsV3 |
| Output | TutorialsV3 |
GET /3/Typeahead/files
Typehead hander for filename completion.
| Input | TypeaheadV3 |
| Output | Schema |
POST /3/UnlockKeys
Unlock all keys in the H2O distributed K/V store, to attempt to recover from a crash.
| Input | UnlockKeysV3 |
| Output | UnlockKeysV3 |
GET /3/WaterMeterCpuTicks/(?.*)
Return a CPU usage snapshot of all cores of all nodes in the H2O cluster.
| Input | WaterMeterCpuTicksV3 |
| Output | WaterMeterCpuTicksV3 |
GET /3/WaterMeterIo
Return IO usage snapshot of all nodes in the H2O cluster.
| Input | WaterMeterIoV3 |
| Output | WaterMeterIoV3 |
GET /3/WaterMeterIo/(?.*)
Return IO usage snapshot of all nodes in the H2O cluster.
| Input | WaterMeterIoV3 |
| Output | WaterMeterIoV3 |
GET /99/Sample
Example of an experimental endpoint. Call via /EXPERIMENTAL/Sample. Experimental endpoints can change at any moment.
| Input | CloudV3 |
| Output | CloudV3 |
REST API Schema Reference
- AboutEntryV3
- AboutV3
- CloudV3
- ClusteringModelBuilderSchema
- ClusteringModelParametersSchema
- ColSpecifierV3
- ColV3
- ColumnSpecsBase
- ConfusionMatrixBase
- ConfusionMatrixV3
- CoxPHModelOutputV3
- CoxPHModelV3
- CoxPHParametersV3
- CoxPHV3
- CreateFrameV3
- DRFGridSearchV3
- DRFModelOutputV3
- DRFModelV3
- DRFParametersV3
- DRFV3
- DStackTraceV3
- DeepLearningModelOutputV3
- DeepLearningModelV3
- DeepLearningParametersV3
- DeepLearningV3
- DownloadDataV3
- EventV3
- ExampleModelOutputV3
- ExampleModelV3
- ExampleParametersV3
- ExampleV3
- FieldMetadataBase
- FieldMetadataV3
- FindV3
- FrameBase
- FrameKeyV3
- FrameSynopsisV3
- FrameV3
- FramesBase
- FramesV3
- GBMGridSearchV3
- GBMModelOutputV3
- GBMModelV3
- GBMParametersV3
- GBMV3
- GLMModelOutputV3
- GLMModelV3
- GLMParametersV3
- GLMV3
- GLRMModelOutputV3
- GLRMModelV3
- GLRMParametersV3
- GLRMV3
- GarbageCollectV3
- GrepModelOutputV3
- GrepModelV3
- GrepParametersV3
- GrepV3
- GridSearchSchema
- H2OErrorV3
- H2OModelBuilderErrorV3
- HeartBeatEvent
- IOEvent
- ImportFilesV3
- InitIDV3
- InteractionV3
- IoStatsEntry
- JStackV3
- JobKeyV3
- JobV3
- JobsV3
- KMeansGridSearchV3
- KMeansModelOutputV3
- KMeansModelV3
- KMeansParametersV3
- KMeansV3
- KeyV3
- KillMinus3V3
- LogAndEchoV3
- LogsV3
- MakeGLMModelV3
- MetadataBase
- MetadataV3
- MissingInserterV3
- ModelBuilderJobV3
- ModelBuilderSchema
- ModelBuildersBase
- ModelBuildersV3
- ModelBuildersVisibilityV3
- ModelKeyV3
- ModelMetricsAutoEncoderV3
- ModelMetricsBase
- ModelMetricsBinomialGLMV3
- ModelMetricsBinomialV3
- ModelMetricsClusteringV3
- ModelMetricsListSchemaV3
- ModelMetricsMultinomialV3
- ModelMetricsPCAV3
- ModelMetricsRegressionGLMV3
- ModelMetricsRegressionV3
- ModelMetricsSVDV3
- ModelOutputSchema
- ModelParameterSchemaV3
- ModelParametersSchema
- ModelSchema
- ModelSynopsisV3
- ModelsBase
- ModelsV3
- NaiveBayesModelOutputV3
- NaiveBayesModelV3
- NaiveBayesParametersV3
- NaiveBayesV3
- NetworkEvent
- NetworkTestV3
- NodePersistentStorageEntryV3
- NodePersistentStorageV3
- NodeV3
- PCAModelOutputV3
- PCAModelV3
- PCAParametersV3
- PCAV3
- ParseSetupV3
- ParseV3
- ProfilerNodeEntryV3
- ProfilerNodeV3
- ProfilerV3
- QuantileParametersV3
- QuantileV3
- RapidsV3
- RemoveAllV3
- RemoveV3
- RouteBase
- RouteV3
- SVDModelOutputV3
- SVDModelV3
- SVDParametersV3
- SVDV3
- Schema
- SchemaMetadataBase
- SchemaMetadataV3
- SharedTreeModelOutputV3
- SharedTreeModelV3
- SharedTreeParametersV3
- SharedTreeV3
- ShutdownV3
- SplitFrameV3
- SupervisedModelBuilderSchema
- SupervisedModelParametersSchema
- SynonymV3
- TimelineV3
- TreeStatsV3
- TutorialsV3
- TwoDimTableBase
- TwoDimTableV3
- TypeaheadV3
- UnlockKeysV3
- ValidationMessageBase
- ValidationMessageV3
- VarImpBase
- VarImpV3
- VecKeyV3
- WaterMeterCpuTicksV3
- WaterMeterIoV3
- Word2VecModelOutputV3
- Word2VecModelV3
- Word2VecParametersV3
- Word2VecV3
AboutEntryV3
namestring | Property name | Out |
valuestring | Property value | Out |
AboutV3
entriesIced[] | List of properties about this running H2O instance | Out |
CloudV3
skip_ticksboolean | skip_ticks | In |
versionstring | version | Out |
node_idxint | Node index number cloud status is collected from (zero-based) | Out |
cloud_namestring | cloud_name | Out |
cloud_sizeint | cloud_size | Out |
cloud_uptime_millislong | cloud_uptime_millis | Out |
cloud_healthyboolean | cloud_healthy | Out |
bad_nodesint | Nodes reporting unhealthy | Out |
consensusboolean | Cloud voting is stable | Out |
lockedboolean | Cloud is accepting new members or not | Out |
nodesIced[] | nodes | Out |
ClusteringModelBuilderSchema
parametersParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
ClusteringModelParametersSchema
kint | Number of clusters | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
ColSpecifierV3
column_namestring | Name of the column | In/Out |
is_member_of_framesstring[] | List of fields which specify columns that must contain this column | In/Out |
ColV3
labelstring | label | Out |
missing_countlong | missing | Out |
zero_countlong | zeros | Out |
positive_infinity_countlong | positive infinities | Out |
negative_infinity_countlong | negative infinities | Out |
minsdouble[] | mins | Out |
maxsdouble[] | maxs | Out |
meandouble | mean | Out |
sigmadouble | sigma | Out |
typestring | datatype: {enum, string, int, real, time, uuid} | Out |
domainstring[] | domain; not-null for enum columns only | Out |
datadouble[] | data | Out |
string_datastring[] | string data | Out |
precisionbyte | decimal precision, -1 for all digits | Out |
histogram_binslong[] | Histogram bins; null if not computed | Out |
histogram_basedouble | Start of histogram bin zero | Out |
histogram_stridedouble | Stride per bin | Out |
percentilesdouble[] | Percentile values, matching the default percentiles | Out |
ColumnSpecsBase
namestring | Column Name | Out |
typestring | Column Type | Out |
formatstring | Column Format (printf) | Out |
descriptionstring | Column Description | Out |
ConfusionMatrixBase
tableTwoDimTable | Annotated confusion matrix | Out |
ConfusionMatrixV3
tableTwoDimTable | Annotated confusion matrix | Out |
CoxPHModelOutputV3
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
CoxPHModelV3
model_idKey | Model key | In/Out |
parametersCoxPHParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputCoxPHOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
CoxPHParametersV3
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
CoxPHV3
parametersCoxPHParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
CreateFrameV3
rowslong | Number of rows | In |
colsint | Number of data columns (in addition to the first response column) | In |
seedlong | Random number seed | In |
randomizeboolean | Whether frame should be randomized | In |
valuelong | Constant value (for randomize=false) | In |
real_rangelong | Range for real variables (-range … range) | In |
categorical_fractiondouble | Fraction of categorical columns (for randomize=true) | In |
factorsint | Factor levels for categorical variables | In |
integer_fractiondouble | Fraction of integer columns (for randomize=true) | In |
integer_rangelong | Range for integer variables (-range … range) | In |
binary_fractiondouble | Fraction of binary columns (for randomize=true) | In |
binary_ones_fractiondouble | Fraction of 1’s in binary columns | In |
missing_fractiondouble | Fraction of missing values | In |
response_factorsint | Number of factor levels of the first column (1=real, 2=binomial, N=multinomial) | In |
has_responseboolean | Whether an additional response column should be generated | In |
keyKey | Job Key | In |
descriptionstring | Job description | In |
destKey | destination key | In/Out |
statusstring | job status | Out |
progressfloat | progress, from 0 to 1 | Out |
progress_msgstring | current progress status description | Out |
start_timelong | Start time | Out |
mseclong | runtime | Out |
exceptionstring | exception | Out |
DRFGridSearchV3
parametersDRFParameters | Basic model builder parameters. | In |
grid_parametersMap | Grid search parameters. | In |
total_modelsint | Number of all models generated by grid search. | Out |
jobJob | Job Key. | Out |
DRFModelOutputV3
variable_importancesTwoDimTable | Variable Importances | Out |
init_fdouble | The Intercept term, the initial model function value to which trees make adjustments | Out |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
DRFModelV3
model_idKey | Model key | In/Out |
parametersDRFParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputDRFOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
DRFParametersV3
mtriesint | Number of columns to randomly select at each level, or -1 for sqrt(#cols) | In |
sample_ratefloat | Sample rate, from 0. to 1.0 | In |
build_tree_one_nodeboolean | Run on one node only; no network overhead but fewer cpus used. Suitable for small datasets. | In |
ntreesint | Number of trees. | In |
max_depthint | Maximum tree depth. | In |
min_rowsint | Fewest allowed observations in a leaf (in R called ‘nodesize’). | In |
nbinsint | Build a histogram of this many bins, then split at the best point | In |
r2_stoppingdouble | Stop making trees when the r^2 metric equals or exceeds this | In |
seedlong | Seed for pseudo random number generator (if applicable) | In |
response_columnVecSpecifier | Response column | In/Out |
balance_classesboolean | Balance training data class counts via over/under-sampling (for imbalanced data). | In/Out |
class_sampling_factorsfloat[] | Desired over/under-sampling ratios per class (in lexicographic order). If not specified, sampling factors will be automatically computed to obtain class balance during training. Requires balance_classes. | In/Out |
max_after_balance_sizefloat | Maximum relative size of the training data after balancing class counts (can be less than 1.0). Requires balance_classes. | In/Out |
max_confusion_matrix_sizeint | Maximum size (# classes) for confusion matrices to be printed in the Logs | In/Out |
max_hit_ratio_kint | Max. number (top K) of predictions to use for hit ratio computation (for multi-class only, 0 to disable) | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
DRFV3
parametersDRFParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
DStackTraceV3
nodestring | Node name | Out |
timelong | Unix epoch time | Out |
thread_tracesstring[] | One trace per thread | Out |
DeepLearningModelOutputV3
weightsKey[] | Frame keys for weight matrices | In |
biasesKey[] | Frame keys for bias vectors | In |
variable_importancesTwoDimTable | Variable Importances | Out |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
DeepLearningModelV3
model_idKey | Model key | In/Out |
parametersDeepLearningParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputDeepLearningModelOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
DeepLearningParametersV3
checkpointKey | Model checkpoint to resume training with | In/Out |
overwrite_with_best_modelboolean | If enabled, override the final model with the best model found during training | In/Out |
autoencoderboolean | Auto-Encoder | In/Out |
use_all_factor_levelsboolean | Use all factor levels of categorical variables. Otherwise, the first factor level is omitted (without loss of accuracy). Useful for variable importances and auto-enabled for autoencoder. | In/Out |
activationenum | Activation function | In/Out |
hiddenint[] | Hidden layer sizes (e.g. 100,100). | In/Out |
epochsdouble | How many times the dataset should be iterated (streamed), can be fractional | In/Out |
train_samples_per_iterationlong | Number of training samples (globally) per MapReduce iteration. Special values are 0: one epoch, -1: all available data (e.g., replicated training data), -2: automatic | In/Out |
target_ratio_comm_to_compdouble | Target ratio of communication overhead to computation. Only for multi-node operation and train_samples_per_iteration=-2 (auto-tuning) | In/Out |
seedlong | Seed for random numbers (affects sampling) - Note: only reproducible when running single threaded | In/Out |
adaptive_rateboolean | Adaptive learning rate | In/Out |
rhodouble | Adaptive learning rate time decay factor (similarity to prior updates) | In/Out |
epsilondouble | Adaptive learning rate smoothing factor (to avoid divisions by zero and allow progress) | In/Out |
ratedouble | Learning rate (higher => less stable, lower => slower convergence) | In/Out |
rate_annealingdouble | Learning rate annealing: rate / (1 + rate_annealing * samples) | In/Out |
rate_decaydouble | Learning rate decay factor between layers (N-th layer: rate*alpha^(N-1)) | In/Out |
momentum_startdouble | Initial momentum at the beginning of training (try 0.5) | In/Out |
momentum_rampdouble | Number of training samples for which momentum increases | In/Out |
momentum_stabledouble | Final momentum after the ramp is over (try 0.99) | In/Out |
nesterov_accelerated_gradientboolean | Use Nesterov accelerated gradient (recommended) | In/Out |
input_dropout_ratiodouble | Input layer dropout ratio (can improve generalization, try 0.1 or 0.2) | In/Out |
hidden_dropout_ratiosdouble[] | Hidden layer dropout ratios (can improve generalization), specify one value per hidden layer, defaults to 0.5 | In/Out |
l1double | L1 regularization (can add stability and improve generalization, causes many weights to become 0) | In/Out |
l2double | L2 regularization (can add stability and improve generalization, causes many weights to be small | In/Out |
max_w2float | Constraint for squared sum of incoming weights per unit (e.g. for Rectifier) | In/Out |
initial_weight_distributionenum | Initial Weight Distribution | In/Out |
initial_weight_scaledouble | Uniform: -value…value, Normal: stddev) | In/Out |
lossenum | Loss function | In/Out |
score_intervaldouble | Shortest time interval (in secs) between model scoring | In/Out |
score_training_sampleslong | Number of training set samples for scoring (0 for all) | In/Out |
score_validation_sampleslong | Number of validation set samples for scoring (0 for all) | In/Out |
score_duty_cycledouble | Maximum duty cycle fraction for scoring (lower: more training, higher: more scoring). | In/Out |
classification_stopdouble | Stopping criterion for classification error fraction on training data (-1 to disable) | In/Out |
regression_stopdouble | Stopping criterion for regression error (MSE) on training data (-1 to disable) | In/Out |
quiet_modeboolean | Enable quiet mode for less output to standard output | In/Out |
score_validation_samplingenum | Method used to sample validation dataset for scoring | In/Out |
diagnosticsboolean | Enable diagnostics for hidden layers | In/Out |
variable_importancesboolean | Compute variable importances for input features (Gedeon method) - can be slow for large networks | In/Out |
fast_modeboolean | Enable fast mode (minor approximation in back-propagation) | In/Out |
force_load_balanceboolean | Force extra load balancing to increase training speed for small datasets (to keep all cores busy) | In/Out |
replicate_training_databoolean | Replicate the entire training dataset onto every node for faster training on small datasets | In/Out |
single_node_modeboolean | Run on a single node for fine-tuning of model parameters | In/Out |
shuffle_training_databoolean | Enable shuffling of training data (recommended if training data is replicated and train_samples_per_iteration is close to #nodes x #rows, of if using balance_classes) | In/Out |
missing_values_handlingenum | Handling of missing values. Either Skip or MeanImputation. | In/Out |
sparseboolean | Sparse data handling (Experimental). | In/Out |
col_majorboolean | Use a column major weight matrix for input layer. Can speed up forward propagation, but might slow down backpropagation (Experimental). | In/Out |
average_activationdouble | Average activation for sparse auto-encoder (Experimental) | In/Out |
sparsity_betadouble | Sparsity regularization (Experimental) | In/Out |
max_categorical_featuresint | Max. number of categorical features, enforced via hashing (Experimental) | In/Out |
reproducibleboolean | Force reproducibility on small data (will be slow - only uses 1 thread) | In/Out |
export_weights_and_biasesboolean | Whether to export Neural Network weights and biases to H2O Frames | In/Out |
response_columnVecSpecifier | Response column | In/Out |
balance_classesboolean | Balance training data class counts via over/under-sampling (for imbalanced data). | In/Out |
class_sampling_factorsfloat[] | Desired over/under-sampling ratios per class (in lexicographic order). If not specified, sampling factors will be automatically computed to obtain class balance during training. Requires balance_classes. | In/Out |
max_after_balance_sizefloat | Maximum relative size of the training data after balancing class counts (can be less than 1.0). Requires balance_classes. | In/Out |
max_confusion_matrix_sizeint | Maximum size (# classes) for confusion matrices to be printed in the Logs | In/Out |
max_hit_ratio_kint | Max. number (top K) of predictions to use for hit ratio computation (for multi-class only, 0 to disable) | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
DeepLearningV3
parametersDeepLearningParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
DownloadDataV3
frame_idKey | Frame to download | In |
hex_stringboolean | Emit double values in a machine readable lossless format with Double.toHexString(). | In |
csvstring | CSV Stream | Out |
filenamestring | Suggested Filename | Out |
EventV3
datestring | Time when the event was recorded. Format is hh:mm:ss:ms | In |
nanoslong | Time in nanos | In |
typeenum | type of recorded event | In |
ExampleModelOutputV3
iterationsint | Iterations executed | In |
maxsdouble[] | (No description available) | In |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
ExampleModelV3
model_idKey | Model key | In/Out |
parametersExampleParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputExampleOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
ExampleParametersV3
max_iterationsint | Maximum training iterations. | In |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
ExampleV3
parametersExampleParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
FieldMetadataBase
schema_namestring | Schema name for this field, if it is_schema, or the name of the enum, if it’s an enum. | In |
namestring | Field name in the Schema | Out |
typestring | Type for this field | Out |
is_schemaboolean | Type for this field is itself a Schema. | Out |
valuePolymorphic | Value for this field | Out |
helpstring | A short help description to appear alongside the field in a UI | Out |
labelstring | The label that should be displayed for the field if the name is insufficient | Out |
requiredboolean | Is this field required, or is the default value generally sufficient? | Out |
levelenum | How important is this field? The web UI uses the level to do a slow reveal of the parameters | Out |
directionenum | Is this field an input, output or inout? | Out |
valuesstring[] | For enum-type fields the allowed values are specified using the values annotation; this is used in UIs to tell the user the allowed values, and for validation | Out |
jsonboolean | Should this field be rendered in the JSON representation? | Out |
is_member_of_framesstring[] | For Vec-type fields this is the set of other Vec-type fields which must contain mutually exclusive values; for example, for a SupervisedModel the response_column must be mutually exclusive with the weights_column | Out |
is_mutually_exclusive_withstring[] | For Vec-type fields this is the set of Frame-type fields which must contain the named column; for example, for a SupervisedModel the response_column must be in both the training_frame and (if it’s set) the validation_frame | Out |
FieldMetadataV3
schema_namestring | Schema name for this field, if it is_schema, or the name of the enum, if it’s an enum. | In |
namestring | Field name in the Schema | Out |
typestring | Type for this field | Out |
is_schemaboolean | Type for this field is itself a Schema. | Out |
valuePolymorphic | Value for this field | Out |
helpstring | A short help description to appear alongside the field in a UI | Out |
labelstring | The label that should be displayed for the field if the name is insufficient | Out |
requiredboolean | Is this field required, or is the default value generally sufficient? | Out |
levelenum | How important is this field? The web UI uses the level to do a slow reveal of the parameters | Out |
directionenum | Is this field an input, output or inout? | Out |
valuesstring[] | For enum-type fields the allowed values are specified using the values annotation; this is used in UIs to tell the user the allowed values, and for validation | Out |
jsonboolean | Should this field be rendered in the JSON representation? | Out |
is_member_of_framesstring[] | For Vec-type fields this is the set of other Vec-type fields which must contain mutually exclusive values; for example, for a SupervisedModel the response_column must be mutually exclusive with the weights_column | Out |
is_mutually_exclusive_withstring[] | For Vec-type fields this is the set of Frame-type fields which must contain the named column; for example, for a SupervisedModel the response_column must be in both the training_frame and (if it’s set) the validation_frame | Out |
FindV3
keyFrame | Frame to search | In |
columnstring | Column, or null for all | In |
rowlong | Starting row for search | In |
matchstring | Value to search for; leave blank for a search for missing values | In |
prevlong | previous row with matching value, or -1 | Out |
nextlong | next row with matching value, or -1 | Out |
FrameBase
frame_idKey | Frame ID | In/Out |
byte_sizelong | Total data size in bytes | Out |
is_textboolean | Is this Frame raw unparsed data? | Out |
FrameKeyV3
namestring | Name (string representation) for this Key. | In/Out |
typestring | Name (string representation) for the type of Keyed this Key points to. | In/Out |
URLstring | URL for the resource that this Key points to, if one exists. | In/Out |
FrameSynopsisV3
frame_idKey | Frame ID | In/Out |
rowslong | Number of rows in the Frame | Out |
columnslong | Number of columns in the Frame | Out |
byte_sizelong | Total data size in bytes | Out |
is_textboolean | Is this Frame raw unparsed data? | Out |
FrameV3
row_offsetlong | Row offset to display | In |
row_countint | Number of rows to display | In/Out |
frame_idKey | Frame ID | In/Out |
checksumlong | checksum | Out |
rowslong | Number of rows in the Frame | Out |
default_percentilesdouble[] | Default percentiles, from 0 to 1 | Out |
columnsVec[] | Columns in the Frame | Out |
compatible_modelsstring[] | Compatible models, if requested | Out |
vec_idsKey | The set of IDs of vectors in the Frame | Out |
chunk_summaryTwoDimTable | Chunk summary | Out |
distribution_summaryTwoDimTable | Distribution summary | Out |
byte_sizelong | Total data size in bytes | Out |
is_textboolean | Is this Frame raw unparsed data? | Out |
FramesBase
frame_idKey | Name of Frame of interest | In |
columnstring | Name of column of interest | In |
find_compatible_modelsboolean | Find and return compatible models? | In |
pathstring | File output path | In |
forceboolean | Overwrite existing file | In |
row_offsetlong | Row offset to display | In/Out |
row_countint | Number of rows to display | In/Out |
framesIced[] | Frames | Out |
compatible_modelsModel[] | Compatible models | Out |
domainstring[][] | Domains | Out |
FramesV3
frame_idKey | Name of Frame of interest | In |
columnstring | Name of column of interest | In |
find_compatible_modelsboolean | Find and return compatible models? | In |
pathstring | File output path | In |
forceboolean | Overwrite existing file | In |
row_offsetlong | Row offset to display | In/Out |
row_countint | Number of rows to display | In/Out |
framesIced[] | Frames | Out |
compatible_modelsModel[] | Compatible models | Out |
domainstring[][] | Domains | Out |
GBMGridSearchV3
parametersGBMParameters | Basic model builder parameters. | In |
grid_parametersMap | Grid search parameters. | In |
total_modelsint | Number of all models generated by grid search. | Out |
jobJob | Job Key. | Out |
GBMModelOutputV3
variable_importancesTwoDimTable | Variable Importances | Out |
init_fdouble | The Intercept term, the initial model function value to which trees make adjustments | Out |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
GBMModelV3
model_idKey | Model key | In/Out |
parametersGBMParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputGBMOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
GBMParametersV3
learn_ratefloat | Learning rate from 0.0 to 1.0 | In |
distributionenum | Distribution function | In |
ntreesint | Number of trees. | In |
max_depthint | Maximum tree depth. | In |
min_rowsint | Fewest allowed observations in a leaf (in R called ‘nodesize’). | In |
nbinsint | Build a histogram of this many bins, then split at the best point | In |
r2_stoppingdouble | Stop making trees when the r^2 metric equals or exceeds this | In |
seedlong | Seed for pseudo random number generator (if applicable) | In |
response_columnVecSpecifier | Response column | In/Out |
balance_classesboolean | Balance training data class counts via over/under-sampling (for imbalanced data). | In/Out |
class_sampling_factorsfloat[] | Desired over/under-sampling ratios per class (in lexicographic order). If not specified, sampling factors will be automatically computed to obtain class balance during training. Requires balance_classes. | In/Out |
max_after_balance_sizefloat | Maximum relative size of the training data after balancing class counts (can be less than 1.0). Requires balance_classes. | In/Out |
max_confusion_matrix_sizeint | Maximum size (# classes) for confusion matrices to be printed in the Logs | In/Out |
max_hit_ratio_kint | Max. number (top K) of predictions to use for hit ratio computation (for multi-class only, 0 to disable) | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
GBMV3
parametersGBMParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
GLMModelOutputV3
coefficients_tableTwoDimTable | Table of Coefficients | In |
standardized_coefficients_magnitudeTwoDimTable | Standardized Coefficient Magnitudes | In |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
GLMModelV3
model_idKey | Model key | In/Out |
parametersGLMParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputGLMOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
GLMParametersV3
familyenum | Family. Use binomial for classification with logistic regression, others are for regression problems. | In |
solverenum | Auto will pick solver better suited for the given dataset, in case of lambda search solvers may be changed during computation. IRLSM is fast on on problems with small number of predictors and for lambda-search with L1 penalty, L_BFGS scales better for datasets with many columns. | In |
alphadouble[] | distribution of regularization between L1 and L2. | In |
lambdadouble[] | regularization strength | In |
lambda_searchboolean | use lambda search starting at lambda max, given lambda is then interpreted as lambda min | In |
nlambdasint | number of lambdas to be used in a search | In |
standardizeboolean | Standardize numeric columns to have zero mean and unit variance | In |
max_iterationsint | Maximum number of iterations | In |
beta_epsilondouble | converge if beta changes less (using L-infinity norm) than beta esilon, ONLY applies to IRLSM solver | In |
objective_epsilondouble | converge if objective value changes less than this | In |
gradient_epsilondouble | converge if objective changes less (using L-infinity norm) than this, ONLY applies to L-BFGS solver | In |
linkenum | (No description available) | In |
interceptboolean | include constant term in the model | In |
priordouble | prior probability for y==1. To be used only for logistic regression iff the data has been sampled and the mean of response does not reflect reality. | In |
lambda_min_ratiodouble | min lambda used in lambda search, specified as a ratio of lambda_max | In |
beta_constraintsKey | beta constraints | In |
max_active_predictorsint | Maximum number of active predictors during computation. Use as a stopping criterium to prevent expensive model building with many predictors. | In |
response_columnVecSpecifier | Response column | In/Out |
balance_classesboolean | Balance training data class counts via over/under-sampling (for imbalanced data). | In/Out |
class_sampling_factorsfloat[] | Desired over/under-sampling ratios per class (in lexicographic order). If not specified, sampling factors will be automatically computed to obtain class balance during training. Requires balance_classes. | In/Out |
max_after_balance_sizefloat | Maximum relative size of the training data after balancing class counts (can be less than 1.0). Requires balance_classes. | In/Out |
max_confusion_matrix_sizeint | Maximum size (# classes) for confusion matrices to be printed in the Logs | In/Out |
max_hit_ratio_kint | Max. number (top K) of predictions to use for hit ratio computation (for multi-class only, 0 to disable) | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
GLMV3
parametersGLMParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
GLRMModelOutputV3
iterationsint | Iterations executed | In |
objectivedouble | Objective value | In |
avg_change_objdouble | Average change in objective value on final iteration | In |
step_sizedouble | Final step size | In |
archetypesdouble[][] | Mapping from training data to lower dimensional k-space | In |
std_deviationdouble[] | Standard deviation of each principal component | In |
eigenvectorsTwoDimTable | Principal components matrix | In |
pc_importanceTwoDimTable | Importance of each principal component | In |
loading_keyKey | Frame key for X matrix | In |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
GLRMModelV3
model_idKey | Model key | In/Out |
parametersGLRMParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputGLRMOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
GLRMParametersV3
transformenum | Transformation of training data | In |
kint | Rank of matrix approximation | In |
lossenum | Loss function | In |
regularization_xenum | Regularization function for X matrix | In |
regularization_yenum | Regularization function for Y matrix | In |
gamma_xdouble | Regularization weight on X matrix | In |
gamma_ydouble | Regularization weight on Y matrix | In |
max_iterationsint | Maximum number of iterations | In |
init_step_sizedouble | Initial step size | In |
min_step_sizedouble | Minimum step size | In |
seedlong | RNG seed for initialization | In |
initenum | Initialization mode | In |
user_pointsKey | User-specified initial Y | In |
loading_keyKey | Frame key to save resulting X | In |
recover_pcaboolean | Recover principal components | In |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
GLRMV3
parametersGLRMParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
GarbageCollectV3
(No fields)
GrepModelOutputV3
matchesstring[] | Matching strings | In |
offsetslong[] | Byte offsets of matches | In |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
GrepModelV3
model_idKey | Model key | In/Out |
parametersGrepParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputGrepOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
GrepParametersV3
regexstring | regex | In |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
GrepV3
parametersGrepParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
GridSearchSchema
parametersParameters | Basic model builder parameters. | In |
grid_parametersMap | Grid search parameters. | In |
total_modelsint | Number of all models generated by grid search. | Out |
jobJob | Job Key. | Out |
H2OErrorV3
timestamplong | Milliseconds since the epoch for the time that this H2OError instance was created. Generally this is a short time since the underlying error ocurred. | Out |
error_urlstring | Error url | Out |
msgstring | Message intended for the end user (a data scientist). | Out |
dev_msgstring | Potentially more detailed message intended for a developer (e.g. a front end engineer or someone designing a language binding). | Out |
http_statusint | HTTP status code for this error. | Out |
valuesMap | Any values that are relevant to reporting or handling this error. Examples are a key name if the error is on a key, or a field name and object name if it’s on a specific field. | Out |
exception_typestring | Exception type, if any. | Out |
exception_msgstring | Raw exception message, if any. | Out |
stacktracestring[] | Stacktrace, if any. | Out |
H2OModelBuilderErrorV3
parametersParameters | Model builder parameters. | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
timestamplong | Milliseconds since the epoch for the time that this H2OError instance was created. Generally this is a short time since the underlying error ocurred. | Out |
error_urlstring | Error url | Out |
msgstring | Message intended for the end user (a data scientist). | Out |
dev_msgstring | Potentially more detailed message intended for a developer (e.g. a front end engineer or someone designing a language binding). | Out |
http_statusint | HTTP status code for this error. | Out |
valuesMap | Any values that are relevant to reporting or handling this error. Examples are a key name if the error is on a key, or a field name and object name if it’s on a specific field. | Out |
exception_typestring | Exception type, if any. | Out |
exception_msgstring | Raw exception message, if any. | Out |
stacktracestring[] | Stacktrace, if any. | Out |
HeartBeatEvent
sendsint | number of sent heartbeats | In |
recvsint | number of received heartbeats | In |
datestring | Time when the event was recorded. Format is hh:mm:ss:ms | In |
nanoslong | Time in nanos | In |
typeenum | type of recorded event | In |
IOEvent
io_flavorstring | flavor of the recorded io (ice/hdfs/…) | In |
nodestring | node where this io event happened | In |
datastring | data info | In |
datestring | Time when the event was recorded. Format is hh:mm:ss:ms | In |
nanoslong | Time in nanos | In |
typeenum | type of recorded event | In |
ImportFilesV3
pathstring | path | In |
filesstring[] | files | Out |
destination_framesstring[] | names | Out |
failsstring[] | fails | Out |
delsstring[] | dels | Out |
InitIDV3
session_keystring | Session ID | Out |
InteractionV3
keyKey | Job Key | In |
descriptionstring | Job description | In |
source_frameKey | Input data frame | In/Out |
factor_columnsstring[] | Factor columns | In/Out |
pairwiseboolean | Whether to create pairwise quadratic interactions between factors (otherwise create one higher-order interaction). Only applicable if there are 3 or more factors. | In/Out |
max_factorsint | Max. number of factor levels in pair-wise interaction terms (if enforced, one extra catch-all factor will be made) | In/Out |
min_occurrenceint | Min. occurrence threshold for factor levels in pair-wise interaction terms | In/Out |
destKey | destination key | In/Out |
statusstring | job status | Out |
progressfloat | progress, from 0 to 1 | Out |
progress_msgstring | current progress status description | Out |
start_timelong | Start time | Out |
mseclong | runtime | Out |
exceptionstring | exception | Out |
IoStatsEntry
backendstring | Back end type | Out |
store_countlong | Number of store events | Out |
store_byteslong | Cumulative stored bytes | Out |
delete_countlong | Number of delete events | Out |
load_countlong | Number of load events | Out |
load_byteslong | Cumulative loaded bytes | Out |
JStackV3
tracesDStackTrace[] | Stacktraces | Out |
JobKeyV3
namestring | Name (string representation) for this Key. | In/Out |
typestring | Name (string representation) for the type of Keyed this Key points to. | In/Out |
URLstring | URL for the resource that this Key points to, if one exists. | In/Out |
JobV3
keyKey | Job Key | In |
descriptionstring | Job description | In |
destKey | destination key | In/Out |
statusstring | job status | Out |
progressfloat | progress, from 0 to 1 | Out |
progress_msgstring | current progress status description | Out |
start_timelong | Start time | Out |
mseclong | runtime | Out |
exceptionstring | exception | Out |
JobsV3
job_idKey | Optional Job identifier | In |
jobsJob[] | jobs | Out |
KMeansGridSearchV3
parametersKMeansParameters | Basic model builder parameters. | In |
grid_parametersMap | Grid search parameters. | In |
total_modelsint | Number of all models generated by grid search. | Out |
jobJob | Job Key. | Out |
KMeansModelOutputV3
centersTwoDimTable | Cluster Centers[k][features] | In |
centers_stdTwoDimTable | Cluster Centers[k][features] on Standardized Data | In |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
KMeansModelV3
model_idKey | Model key | In/Out |
parametersKMeansParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputKMeansOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
KMeansParametersV3
user_pointsKey | User-specified points | In |
max_iterationsint | Maximum training iterations | In |
standardizeboolean | Standardize columns | In |
seedlong | RNG Seed | In |
initenum | Initialization mode | In |
kint | Number of clusters | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
KMeansV3
parametersKMeansParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
KeyV3
namestring | Name (string representation) for this Key. | In/Out |
typestring | Name (string representation) for the type of Keyed this Key points to. | In/Out |
URLstring | URL for the resource that this Key points to, if one exists. | In/Out |
KillMinus3V3
(No fields)
LogAndEchoV3
messagestring | Message to be Logged and Echoed | In |
LogsV3
nodeidxint | Index of node to query ticks for (0-based). -1 means current node. | In |
namestring | Which specific log file to read from the log file directory. If left unspecified, the system chooses a default for you. | In |
logstring | Content of log file | Out |
MakeGLMModelV3
modelKey | source model | In |
destKey | destination key | In |
namesstring[] | coefficient names | In |
betadouble[] | new glm coefficients | In |
thresholdfloat | decision threshold for label-generation | In |
MetadataBase
numint | Number for specifying an endpoint | In |
http_methodstring | HTTP method (GET, POST, DELETE) if fetching by path | In |
pathstring | Path for specifying an endpoint | In |
classnamestring | Class name, for fetching docs for a schema (DEPRECATED) | In |
schemanamestring | Schema name (e.g., DocsV1), for fetching docs for a schema | In |
routesRoute[] | List of endpoint routes | Out |
schemasSchemaMetadata[] | List of schemas | Out |
markdownstring | Table of Contents Markdown | Out |
MetadataV3
numint | Number for specifying an endpoint | In |
http_methodstring | HTTP method (GET, POST, DELETE) if fetching by path | In |
pathstring | Path for specifying an endpoint | In |
classnamestring | Class name, for fetching docs for a schema (DEPRECATED) | In |
schemanamestring | Schema name (e.g., DocsV1), for fetching docs for a schema | In |
routesRoute[] | List of endpoint routes | Out |
schemasSchemaMetadata[] | List of schemas | Out |
markdownstring | Table of Contents Markdown | Out |
MissingInserterV3
datasetKey | dataset | In |
fractiondouble | Fraction of data to replace with a missing value | In |
seedlong | Seed | In |
keyKey | Job Key | In |
descriptionstring | Job description | In |
destKey | destination key | In/Out |
statusstring | job status | Out |
progressfloat | progress, from 0 to 1 | Out |
progress_msgstring | current progress status description | Out |
start_timelong | Start time | Out |
mseclong | runtime | Out |
exceptionstring | exception | Out |
ModelBuilderJobV3
keyKey | Job Key | In |
descriptionstring | Job description | In |
destKey | destination key | In/Out |
parametersParameters | Model builder parameters. | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
statusstring | job status | Out |
progressfloat | progress, from 0 to 1 | Out |
progress_msgstring | current progress status description | Out |
start_timelong | Start time | Out |
mseclong | runtime | Out |
exceptionstring | exception | Out |
ModelBuilderSchema
parametersParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
ModelBuildersBase
algostring | Algo of ModelBuilder of interest | In |
model_buildersMap | ModelBuilders | Out |
ModelBuildersV3
algostring | Algo of ModelBuilder of interest | In |
model_buildersMap | ModelBuilders | Out |
ModelBuildersVisibilityV3
valuestring | Stable, Beta, Experimental | In/Out |
ModelKeyV3
namestring | Name (string representation) for this Key. | In/Out |
typestring | Name (string representation) for the type of Keyed this Key points to. | In/Out |
URLstring | URL for the resource that this Key points to, if one exists. | In/Out |
ModelMetricsAutoEncoderV3
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsBase
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsBinomialGLMV3
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
residual_deviancedouble | residual deviance | Out |
null_deviancedouble | null deviance | Out |
AICdouble | AIC | Out |
null_degrees_of_freedomlong | null DOF | Out |
residual_degrees_of_freedomlong | residual DOF | Out |
r2double | The R^2 for this scoring run. | Out |
loglossdouble | The logarithmic loss for this scoring run. | Out |
AUCdouble | The AUC for this scoring run. | Out |
Ginidouble | The Gini score for this scoring run. | Out |
domainstring[] | The class labels of the response. | Out |
thresholds_and_metric_scoresTwoDimTable | The Metrics for various thresholds. | Out |
max_criteria_and_metric_scoresTwoDimTable | The Metrics for various criteria. | Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsBinomialV3
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
r2double | The R^2 for this scoring run. | Out |
loglossdouble | The logarithmic loss for this scoring run. | Out |
AUCdouble | The AUC for this scoring run. | Out |
Ginidouble | The Gini score for this scoring run. | Out |
domainstring[] | The class labels of the response. | Out |
thresholds_and_metric_scoresTwoDimTable | The Metrics for various thresholds. | Out |
max_criteria_and_metric_scoresTwoDimTable | The Metrics for various criteria. | Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsClusteringV3
tot_withinssdouble | Within Cluster Sum of Square Error | In |
totssdouble | Total Sum of Square Error to Grand Mean | In |
betweenssdouble | Between Cluster Sum of Square Error | In |
centroid_statsTwoDimTable | Centroid Statistics | In |
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsListSchemaV3
modelKey | Key of Model of interest (optional) | In |
frameKey | Key of Frame of interest (optional) | In |
reconstruction_errorboolean | Compute reconstruction error (optional, only for Deep Learning AutoEncoder models) | In |
deep_features_hidden_layerint | Extract Deep Features for given hidden layer (optional, only for Deep Learning models) | In |
predictions_frameKey | Key of predictions frame, if predictions are requested (optional) | In/Out |
model_metricsModelMetrics[] | ModelMetrics | Out |
ModelMetricsMultinomialV3
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
r2double | The R^2 for this scoring run. | Out |
hit_ratio_tableTwoDimTable | The hit ratio table for this scoring run. | Out |
cmConfusionMatrix | The ConfusionMatrix object for this scoring run. | Out |
loglossdouble | The logarithmic loss for this scoring run. | Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsPCAV3
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsRegressionGLMV3
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
residual_deviancedouble | residual deviance | Out |
null_deviancedouble | null deviance | Out |
AICdouble | AIC | Out |
null_degrees_of_freedomlong | null DOF | Out |
residual_degrees_of_freedomlong | residual DOF | Out |
r2double | The R^2 for this scoring run. | Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsRegressionV3
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
r2double | The R^2 for this scoring run. | Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelMetricsSVDV3
modelKey | The model used for this scoring run. | In/Out |
model_checksumlong | The checksum for the model used for this scoring run. | In/Out |
frameKey | The frame used for this scoring run. | In/Out |
frame_checksumlong | The checksum for the frame used for this scoring run. | In/Out |
descriptionstring | Optional description for this scoring run (to note out-of-bag, sampled data, etc.) | Out |
model_categoryenum | The category (e.g., Clustering) for the model used for this scoring run. | Out |
scoring_timelong | The time in mS since the epoch for the start of this scoring run. | Out |
predictionsFrame | Predictions Frame. | Out |
MSEdouble | The Mean Squared Error of the prediction for this scoring run. | Out |
ModelOutputSchema
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
ModelParameterSchemaV3
is_member_of_framesstring[] | For Vec-type fields this is the set of other Vec-type fields which must contain mutually exclusive values; for example, for a SupervisedModel the response_column must be mutually exclusive with the weights_column | In |
is_mutually_exclusive_withstring[] | For Vec-type fields this is the set of Frame-type fields which must contain the named column; for example, for a SupervisedModel the response_column must be in both the training_frame and (if it’s set) the validation_frame | In |
namestring | name in the JSON, e.g. “lambda” | Out |
labelstring | label in the UI, e.g. “lambda” | Out |
helpstring | help for the UI, e.g. “regularization multiplier, typically used for foo bar baz etc.” | Out |
requiredboolean | the field is required | Out |
typestring | Java type, e.g. “double” | Out |
default_valuePolymorphic | default value, e.g. 1 | Out |
actual_valuePolymorphic | actual value as set by the user and / or modified by the ModelBuilder, e.g., 10 | Out |
levelstring | the importance of the parameter, used by the UI, e.g. “critical”, “extended” or “expert” | Out |
valuesstring[] | list of valid values for use by the front-end | Out |
ModelParametersSchema
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
ModelSchema
model_idKey | Model key | In/Out |
parametersParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
ModelSynopsisV3
model_idKey | Model key | In/Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
ModelsBase
model_idKey | Name of Model of interest | In |
previewboolean | Return potentially abridged model suitable for viewing in a browser | In |
find_compatible_framesboolean | Find and return compatible frames? | In |
modelsModel[] | Models | Out |
compatible_framesFrame[] | Compatible frames | Out |
ModelsV3
model_idKey | Name of Model of interest | In |
previewboolean | Return potentially abridged model suitable for viewing in a browser | In |
find_compatible_framesboolean | Find and return compatible frames? | In |
modelsModel[] | Models | Out |
compatible_framesFrame[] | Compatible frames | Out |
NaiveBayesModelOutputV3
levelsstring[] | Categorical levels of the response | In |
aprioriTwoDimTable | A-priori probabilities of the response | In |
pcondTwoDimTable[] | Conditional probabilities of the predictors | In |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
NaiveBayesModelV3
model_idKey | Model key | In/Out |
parametersNaiveBayesParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputNaiveBayesOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
NaiveBayesParametersV3
laplacedouble | Laplace smoothing parameter | In |
min_sdevdouble | Min. standard deviation to use for observations with not enough data | In |
eps_sdevdouble | Cutoff below which standard deviation is replaced with min_sdev | In |
min_probdouble | Min. probability to use for observations with not enough data | In |
eps_probdouble | Cutoff below which probability is replaced with min_prob | In |
compute_metricsboolean | Compute metrics on training data | In |
response_columnVecSpecifier | Response column | In/Out |
balance_classesboolean | Balance training data class counts via over/under-sampling (for imbalanced data). | In/Out |
class_sampling_factorsfloat[] | Desired over/under-sampling ratios per class (in lexicographic order). If not specified, sampling factors will be automatically computed to obtain class balance during training. Requires balance_classes. | In/Out |
max_after_balance_sizefloat | Maximum relative size of the training data after balancing class counts (can be less than 1.0). Requires balance_classes. | In/Out |
max_confusion_matrix_sizeint | Maximum size (# classes) for confusion matrices to be printed in the Logs | In/Out |
max_hit_ratio_kint | Max. number (top K) of predictions to use for hit ratio computation (for multi-class only, 0 to disable) | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
NaiveBayesV3
parametersNaiveBayesParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
NetworkEvent
is_sendboolean | Boolean flag distinguishing between sends (true) and receives(false) | In |
protocolstring | network protocol (UDP/TCP) | In |
msg_typestring | UDP type (exec,ack, ackack,… | In |
fromstring | Sending node | In |
tostring | Receiving node | In |
datastring | Pretty print of the first few bytes of the msg payload. Contains class name for tasks. | In |
datestring | Time when the event was recorded. Format is hh:mm:ss:ms | In |
nanoslong | Time in nanos | In |
typeenum | type of recorded event | In |
NetworkTestV3
microseconds_collectivedouble[] | Collective broadcast/reduce times in microseconds (for each message size) | Out |
bandwidths_collectivedouble[] | Collective bandwidths in Bytes/sec (for each message size, for each node) | Out |
microsecondsdouble[][] | Round-trip times in microseconds (for each message size, for each node) | Out |
bandwidthsdouble[][] | Bi-directional bandwidths in Bytes/sec (for each message size, for each node) | Out |
nodesstring[] | Nodes | Out |
tableTwoDimTable | NetworkTestResults | Out |
NodePersistentStorageEntryV3
categorystring | Category name | Out |
namestring | Key name | Out |
sizelong | Size in bytes of value | Out |
timestamp_millislong | Epoch time in milliseconds of when the value was written | Out |
NodePersistentStorageV3
categorystring | Category name | In/Out |
namestring | Key name | In/Out |
valuestring | Value | In/Out |
configuredboolean | Configured | Out |
existsboolean | Exists | Out |
entriesIced[] | List of entries | Out |
NodeV3
h2ostring | IP | Out |
ip_portstring | IP address and port in the form a.b.c.d:e | Out |
healthyboolean | (now-last_ping)<HeartbeatThread.TIMEOUT | Out |
last_pinglong | Time (in msec) of last ping | Out |
sys_loadfloat | System load; average #runnables/#cores | Out |
gflopsdouble | Linpack GFlops | Out |
mem_bwdouble | Memory Bandwidth | Out |
total_value_sizelong | Data on Node (memory or disk) | Out |
mem_value_sizelong | Data on Node (memory only) | Out |
num_keysint | id="local-keys">local keys< | Out |
free_memlong | Free heap | Out |
tot_memlong | Total heap | Out |
max_memlong | Max heap | Out |
free_disklong | Free disk | Out |
max_disklong | Max disk | Out |
rpcs_activeint | Active Remote Procedure Calls | Out |
fjthrdsshort[] | F/J Thread count, by priority | Out |
fjqueueshort[] | F/J Task count, by priority | Out |
tcps_activeint | Open TCP connections | Out |
open_fdsint | Open File Descripters | Out |
num_cpusint | num_cpus | Out |
cpus_allowedint | cpus_allowed | Out |
nthreadsint | nthreads | Out |
my_cpu_pctint | System CPU percentage used by this H2O process in last interval | Out |
sys_cpu_pctint | System CPU percentage used by everything in last interval | Out |
pidstring | PID | Out |
PCAModelOutputV3
std_deviationdouble[] | Standard deviation of each principal component | In |
eigenvectorsTwoDimTable | Principal components matrix | In |
pc_importanceTwoDimTable | Importance of each principal component | In |
loading_keyKey | Frame key for loading matrix | In |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
PCAModelV3
model_idKey | Model key | In/Out |
parametersPCAParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputPCAOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
PCAParametersV3
transformenum | Transformation of training data | In |
kint | Rank of matrix approximation | In |
max_iterationsint | Maximum training iterations | In |
seedlong | RNG seed for initialization | In |
loading_keyKey | Frame key to save resulting X | In |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
PCAV3
parametersPCAParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
ParseSetupV3
source_framesKey[] | Source frames | In/Out |
parse_typeenum | Parser type | In/Out |
separatorbyte | Field separator | In/Out |
single_quotesboolean | Single quotes | In/Out |
check_headerint | Check header: 0 means guess, +1 means 1st line is header not data, -1 means 1st line is data not header | In/Out |
column_namesstring[] | Column names | In/Out |
column_typesstring[] | Value types for columns | In/Out |
na_stringsstring[][] | NA strings for columns | In/Out |
destination_framestring | Suggested name | Out |
header_lineslong | Number of header lines found | Out |
number_columnsint | Number of columns | Out |
datastring[][] | Sample data | Out |
chunk_sizeint | Size of individual parse tasks | Out |
ParseV3
destination_frameKey | Final frame name | In |
source_framesKey[] | Source frames | In |
parse_typeenum | Parser type | In |
separatorbyte | Field separator | In |
single_quotesboolean | Single Quotes | In |
check_headerint | Check header: 0 means guess, +1 means 1st line is header not data, -1 means 1st line is data not header | In |
number_columnsint | Number of columns | In |
column_namesstring[] | Column names | In |
column_typesstring[] | Value types for columns | In |
domainsstring[][] | Domains for categorical columns | In |
na_stringsstring[][] | NA strings for columns | In |
chunk_sizeint | Size of individual parse tasks | In |
delete_on_doneboolean | Delete input key after parse | In |
blockingboolean | Block until the parse completes (as opposed to returning early and requiring polling | In |
remove_frameboolean | Remove frame after blocking parse, and return array of Vecs | In |
jobJob | Parse job | Out |
rowslong | Rows | Out |
vec_idsKey | Vec IDs | Out |
ProfilerNodeEntryV3
stacktracestring | Stack trace | Out |
countint | Profile Count | Out |
ProfilerNodeV3
node_namestring | Node names | Out |
timestamplong | Timestamp (millis since epoch) | Out |
entriesIced[] | Profile entry list | Out |
ProfilerV3
depthint | Stack trace depth | In |
nodesIced[] | (No description available) | Out |
QuantileParametersV3
probsdouble[] | Probabilities for quantiles | In |
combine_methodenum | How to combine quantiles for even sample sizes | In |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
QuantileV3
parametersQuantileParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
RapidsV3
aststring | An Abstract Syntax Tree. | In |
funstring | An array of function definitions. | In |
ast_keyKey | A pointer to a Frame | In |
errorstring | Parsing error, if any | Out |
keyKey | Result key | Out |
num_rowslong | Rows in Frame result | Out |
num_colsint | Columns in Frame result | Out |
scalardouble | Scalar result | Out |
funstrstring | Function result | Out |
col_namesstring[] | Column Names | Out |
stringstring | String result | Out |
resultstring | result | Out |
evaluatedboolean | Was evaluated | Out |
headstring[][] | Head of a Frame result | Out |
result_typeint | Result Type. | Out |
vec_idsKey | Vec keys for key result | Out |
RemoveAllV3
(No fields)
RemoveV3
keyKey | Object to be removed. | In |
RouteBase
http_methodstring | (No description available) | Out |
url_patternstring | (No description available) | Out |
summarystring | (No description available) | Out |
handler_classstring | (No description available) | Out |
handler_methodstring | (No description available) | Out |
input_schemastring | (No description available) | Out |
output_schemastring | (No description available) | Out |
doc_methodstring | (No description available) | Out |
path_paramsstring[] | (No description available) | Out |
markdownstring | (No description available) | Out |
RouteV3
http_methodstring | (No description available) | Out |
url_patternstring | (No description available) | Out |
summarystring | (No description available) | Out |
handler_classstring | (No description available) | Out |
handler_methodstring | (No description available) | Out |
input_schemastring | (No description available) | Out |
output_schemastring | (No description available) | Out |
doc_methodstring | (No description available) | Out |
path_paramsstring[] | (No description available) | Out |
markdownstring | (No description available) | Out |
SVDModelOutputV3
vdouble[][] | Right singular vectors | In |
ddouble[] | Singular values | In |
u_keyKey | Frame key of left singular vectors | In |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
SVDModelV3
model_idKey | Model key | In/Out |
parametersSVDParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputSVDOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
SVDParametersV3
transformenum | Transformation of training data | In |
nvint | Number of right singular vectors | In |
max_iterationsint | Maximum iterations | In |
seedlong | RNG seed for k-means++ initialization | In |
keep_uboolean | Save left singular vectors? | In |
u_keyKey | Frame key to save left singular vectors | In |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
SVDV3
parametersSVDParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
Schema
(No fields)
SchemaMetadataBase
versionint | Version number of the Schema. | In |
namestring | Simple name of the Schema. NOTE: the schema_names form a single namespace. | In |
superclassstring | Simple name of the superclass of the Schema. NOTE: the schema_names form a single namespace. | In |
typestring | Simple name of H2O type that this Schema represents. Must not be changed after creation (treat as final). | In |
fieldsFieldMetadata[] | All the public fields of the schema | Out |
markdownstring | Documentation for the schema in Markdown format with GitHub extensions | Out |
SchemaMetadataV3
versionint | Version number of the Schema. | In |
namestring | Simple name of the Schema. NOTE: the schema_names form a single namespace. | In |
superclassstring | Simple name of the superclass of the Schema. NOTE: the schema_names form a single namespace. | In |
typestring | Simple name of H2O type that this Schema represents. Must not be changed after creation (treat as final). | In |
fieldsFieldMetadata[] | All the public fields of the schema | Out |
markdownstring | Documentation for the schema in Markdown format with GitHub extensions | Out |
SharedTreeModelOutputV3
variable_importancesTwoDimTable | Variable Importances | Out |
init_fdouble | The Intercept term, the initial model function value to which trees make adjustments | Out |
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
SharedTreeModelV3
model_idKey | Model key | In/Out |
parametersParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
SharedTreeParametersV3
ntreesint | Number of trees. | In |
max_depthint | Maximum tree depth. | In |
min_rowsint | Fewest allowed observations in a leaf (in R called ‘nodesize’). | In |
nbinsint | Build a histogram of this many bins, then split at the best point | In |
r2_stoppingdouble | Stop making trees when the r^2 metric equals or exceeds this | In |
seedlong | Seed for pseudo random number generator (if applicable) | In |
response_columnVecSpecifier | Response column | In/Out |
balance_classesboolean | Balance training data class counts via over/under-sampling (for imbalanced data). | In/Out |
class_sampling_factorsfloat[] | Desired over/under-sampling ratios per class (in lexicographic order). If not specified, sampling factors will be automatically computed to obtain class balance during training. Requires balance_classes. | In/Out |
max_after_balance_sizefloat | Maximum relative size of the training data after balancing class counts (can be less than 1.0). Requires balance_classes. | In/Out |
max_confusion_matrix_sizeint | Maximum size (# classes) for confusion matrices to be printed in the Logs | In/Out |
max_hit_ratio_kint | Max. number (top K) of predictions to use for hit ratio computation (for multi-class only, 0 to disable) | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
SharedTreeV3
parametersParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
ShutdownV3
(No fields)
SplitFrameV3
datasetKey | Dataset | In |
ratiosdouble[] | Split ratios - resulting number of split is ratios.length+1 | In |
keyKey | Job Key | In |
descriptionstring | Job description | In |
destination_framesKey[] | Destination keys for each output frame split. | In/Out |
destKey | destination key | In/Out |
statusstring | job status | Out |
progressfloat | progress, from 0 to 1 | Out |
progress_msgstring | current progress status description | Out |
start_timelong | Start time | Out |
mseclong | runtime | Out |
exceptionstring | exception | Out |
SupervisedModelBuilderSchema
parametersParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |
SupervisedModelParametersSchema
response_columnVecSpecifier | Response column | In/Out |
balance_classesboolean | Balance training data class counts via over/under-sampling (for imbalanced data). | In/Out |
class_sampling_factorsfloat[] | Desired over/under-sampling ratios per class (in lexicographic order). If not specified, sampling factors will be automatically computed to obtain class balance during training. Requires balance_classes. | In/Out |
max_after_balance_sizefloat | Maximum relative size of the training data after balancing class counts (can be less than 1.0). Requires balance_classes. | In/Out |
max_confusion_matrix_sizeint | Maximum size (# classes) for confusion matrices to be printed in the Logs | In/Out |
max_hit_ratio_kint | Max. number (top K) of predictions to use for hit ratio computation (for multi-class only, 0 to disable) | In/Out |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
SynonymV3
keyKey | A word2vec model key. | In |
targetstring | The target string to find synonyms. | In |
cntint | Find the top cnt synonyms of the target word. | In |
synonymsstring[] | The synonyms. | Out |
cos_simfloat[] | The cosine similarities. | Out |
TimelineV3
nowlong | Current time in millis. | Out |
selfstring | This node | Out |
eventsIced[] | recorded timeline events | Out |
TreeStatsV3
min_depthint | minDepth | In |
max_depthint | maxDepth | In |
mean_depthfloat | meanDepth | In |
min_leavesint | minLeaves | In |
max_leavesint | maxLeaves | In |
mean_leavesfloat | meanLeaves | In |
TutorialsV3
(No fields)
TwoDimTableBase
namestring | Table Name | Out |
descriptionstring | Table Description | Out |
columnsIced[] | Column Specification | Out |
rowcountint | Number of Rows | Out |
dataPolymorphic[][] | Table Data (col-major) | Out |
TwoDimTableV3
namestring | Table Name | Out |
descriptionstring | Table Description | Out |
columnsIced[] | Column Specification | Out |
rowcountint | Number of Rows | Out |
dataPolymorphic[][] | Table Data (col-major) | Out |
TypeaheadV3
srcstring | training_frame | In |
limitint | limit | In |
matchesstring[] | matches | Out |
UnlockKeysV3
(No fields)
ValidationMessageBase
message_typestring | Type of validation message (ERROR, WARN, INFO, HIDE) | Out |
field_namestring | Field to which the message applies | Out |
messagestring | Message text | Out |
ValidationMessageV3
message_typestring | Type of validation message (ERROR, WARN, INFO, HIDE) | Out |
field_namestring | Field to which the message applies | Out |
messagestring | Message text | Out |
VarImpBase
varimpfloat[] | Variable importance of individual variables | Out |
namesstring[] | Names of variables | Out |
VarImpV3
varimpfloat[] | Variable importance of individual variables | Out |
namesstring[] | Names of variables | Out |
VecKeyV3
namestring | Name (string representation) for this Key. | In/Out |
typestring | Name (string representation) for the type of Keyed this Key points to. | In/Out |
URLstring | URL for the resource that this Key points to, if one exists. | In/Out |
WaterMeterCpuTicksV3
nodeidxint | Index of node to query ticks for (0-based) | In |
cpu_tickslong[][] | array of tick counts per core | Out |
WaterMeterIoV3
nodeidxint | Index of node to query ticks for (0-based) | In |
persist_statsIced[] | array of IO info | Out |
Word2VecModelOutputV3
namesstring[] | Column names. | Out |
domainsstring[][] | Domains for categorical (enum) columns. | Out |
model_categoryenum | Category of the model (e.g., Binomial). | Out |
model_summaryTwoDimTable | Model summary | Out |
scoring_historyTwoDimTable | Scoring history | Out |
training_metricsModelMetrics | Training data model metrics | Out |
validation_metricsModelMetrics | Validation data model metrics | Out |
helpMap | Help information for output fields | Out |
Word2VecModelV3
model_idKey | Model key | In/Out |
parametersWord2VecParameters | The build parameters for the model (e.g. K for KMeans). | Out |
outputWord2VecOutput | The build output for the model (e.g. the cluster centers for KMeans). | Out |
compatible_framesstring[] | Compatible frames, if requested | Out |
checksumlong | Checksum for all the things that go into building the Model. | Out |
algostring | The algo name for this Model. | Out |
algo_full_namestring | The pretty algo name for this Model (e.g., Generalized Linear Model, rather than GLM). | Out |
Word2VecParametersV3
vecSizeint | Set size of word vectors | In |
windowSizeint | Set max skip length between words | In |
sentSampleRatefloat | Set threshold for occurrence of words. Those that appear with higher frequency in the training data will be randomly down-sampled; useful range is (0, 1e-5) | In |
normModelenum | Use Hierarchical Softmax or Negative Sampling | In |
negSampleCntint | Number of negative examples, common values are 3 - 10 (0 = not used) | In |
epochsint | Number of training iterations to run | In |
minWordFreqint | This will discard words that appear less than | In |
initLearningRatefloat | Set the starting learning rate | In |
wordModelenum | Use the continuous bag of words model or the Skip-Gram model | In |
model_idKey | Destination id for this model; auto-generated if not specified | In/Out |
training_frameKey | Training frame | In/Out |
validation_frameKey | Validation frame | In/Out |
ignored_columnsstring[] | Ignored columns | In/Out |
ignore_const_colsboolean | Ignore constant columns | In/Out |
score_each_iterationboolean | Whether to score during each iteration of model training | In/Out |
Word2VecV3
parametersWord2VecParameters | Model builder parameters. | In |
__http_statusint | HTTP status to return for this build. | In |
algostring | The algo name for this ModelBuilder. | Out |
algo_full_namestring | The pretty algo name for this ModelBuilder (e.g., Generalized Linear Model, rather than GLM). | Out |
can_buildenum[] | Model categories this ModelBuilder can build. | Out |
visibilityenum | Should the builder always be visible, be marked as beta, or only visible if the user starts up with the experimental flag? | Out |
jobJob | Job Key | Out |
messagesValidationMessage[] | Parameter validation messages | Out |
error_countint | Count of parameter validation errors | Out |